Decoding RNA Biology: A Comprehensive Guide to RBP Interactions with lncRNAs and miRNAs
This article provides a systematic overview of the intricate regulatory networks formed by RNA-binding proteins (RBPs) with non-coding RNAs, specifically long non-coding RNAs (lncRNAs) and microRNAs (miRNAs).
Decoding RNA Biology: A Comprehensive Guide to RBP Interactions with lncRNAs and miRNAs
Abstract
This article provides a systematic overview of the intricate regulatory networks formed by RNA-binding proteins (RBPs) with non-coding RNAs, specifically long non-coding RNAs (lncRNAs) and microRNAs (miRNAs). Targeting researchers and drug development professionals, we explore the foundational principles of these interactions, detailing state-of-the-art methodologies for their identification and characterization. We address common experimental challenges and optimization strategies, and compare validation techniques for confirming functional outcomes. The synthesis offers a roadmap for leveraging these interactions in understanding disease mechanisms and developing novel therapeutic modalities.
Unveiling the Regulatory Nexus: Core Principles of RBP-ncRNA Interactions
Within the complex landscape of gene regulation, the dynamic interactions between RNA-binding proteins (RBPs), long non-coding RNAs (lncRNAs), and microRNAs (miRNAs) form a critical nexus. This overview details the core definitions, functions, and quantitative profiles of these key molecular players, framed within the context of their intricate cross-talk in cellular homeostasis and disease. Understanding these interactions is fundamental to advancing therapeutic interventions.
Defining the Core Players
RNA-Binding Proteins (RBPs)
RBPs are proteins that associate with single or double-stranded RNA through specialized RNA-binding domains (RBDs). They govern every aspect of an RNA's lifecycle, from splicing, transport, and stability to translation.
Common RBDs and Functions:
- RRM (RNA Recognition Motif): The most abundant domain; recognizes 2-8 nucleotide sequences.
- KH (K-homology) Domain: Binds single-stranded RNA/DNA.
- Zinc Finger Motifs: Often involved in transcriptional regulation but some bind RNA.
- DEAD-box Helicase Domain: Involved in ATP-dependent RNA unwinding.
Long Non-Coding RNAs (lncRNAs)
lncRNAs are a diverse class of transcripts >200 nucleotides with limited or no protein-coding potential. They function as scaffolds, guides, decoys, or signals to modulate gene expression at epigenetic, transcriptional, and post-transcriptional levels.
MicroRNAs (miRNAs)
miRNAs are small (~22 nucleotide), highly conserved non-coding RNAs that primarily regulate gene expression post-transcriptionally. They guide the RNA-induced silencing complex (RISC) to target mRNAs via base-pairing, leading to mRNA degradation or translational repression.
Table 1: Comparative Overview of RBPs, lncRNAs, and miRNAs
| Feature | RNA-Binding Proteins (RBPs) | Long Non-Coding RNAs (lncRNAs) | MicroRNAs (miRNAs) |
|---|---|---|---|
| Chemical Nature | Protein | RNA (transcript >200 nt) | RNA (mature form ~22 nt) |
| Estimated Count in Human Genome | ~1,500 - 2,000 | >17,000 loci | ~2,300 mature sequences |
| Primary Function | Regulate RNA metabolism & function | Multifunctional regulators (scaffold, guide, decoy) | Post-transcriptional gene silencing |
| Key Mechanism | Bind via structured domains; enzymatic/modification activity | Sequence-specific binding & structural interactions | Imperfect base-pairing with mRNA 3'UTR |
| Typical Target | All RNA classes (mRNA, lncRNA, miRNA, etc.) | Chromatin, RBPs, transcription factors, miRNAs | Complementary mRNA sequences |
| Dysregulation Implicated In | Cancer, neurodegeneration, metabolic disorders | Cancer, cardiovascular disease, neurological disorders | Cancer, viral infections, immune disorders |
Data synthesized from recent reviews in *Nature Reviews Genetics and Nucleic Acids Research (2023-2024).*
Key Methodologies for Studying Interactions
To dissect the interactions within the RBP-lncRNA-miRNA axis, researchers employ a suite of advanced techniques.
Protocol: Cross-Linking and Immunoprecipitation (CLIP-seq)
Purpose: To map genome-wide binding sites of an RBP on its RNA targets (including lncRNAs and pre-miRNAs). Detailed Workflow:
- In Vivo Crosslinking: Cells are irradiated with UV-C light (254 nm) to create covalent bonds between the RBP and its directly bound RNA.
- Cell Lysis and Immunoprecipitation: Cells are lysed, and the RBP-RNA complexes are isolated using antibodies specific to the RBP of interest.
- RNA Processing: Proteins are digested, and the bound RNA is recovered, reverse-transcribed, and converted into a sequencing library.
- High-Throughput Sequencing & Analysis: Libraries are sequenced. Bioinformatics tools (e.g.,
CLIPper,Piranha) identify significant binding peaks.
Protocol: miRNA Target Identification (AGO-PAR-CLIP)
Purpose: To identify miRNA binding sites by capturing RNAs bound to Argonaute (AGO), the core RBP component of RISC. Detailed Workflow:
- Incorporation of Photoactivatable Ribonucleoside: Cells are cultured with 4-thiouridine (4SU), which incorporates into nascent RNA.
- Crosslinking: UV light at 365 nm crosslinks 4SU-labeled RNA to interacting AGO proteins.
- Immunoprecipitation and Sequencing: AGO-RNA complexes are isolated, and the RNA is prepared for sequencing. T-to-C mutations in the reads mark the precise crosslinked nucleotide, revealing the miRNA binding site.
Protocol: Functional lncRNA Screening (CRISPRi)
Purpose: To interrogate lncRNA function in a high-throughput manner. Detailed Workflow:
- Library Design: A sgRNA library is designed to target transcription start sites or regulatory elements of thousands of lncRNAs.
- Viral Transduction: A cell line stably expressing dCas9-KRAB (a transcriptional repressor) is transduced with the sgRNA library.
- Phenotypic Selection: Cells are subjected to a selective pressure (e.g., drug treatment, proliferation). Surviving cells are collected.
- Sequencing & Hit Identification: sgRNA abundance in pre- and post-selection pools is quantified by sequencing to identify lncRNAs essential for the phenotype.
Diagrammatic Representations
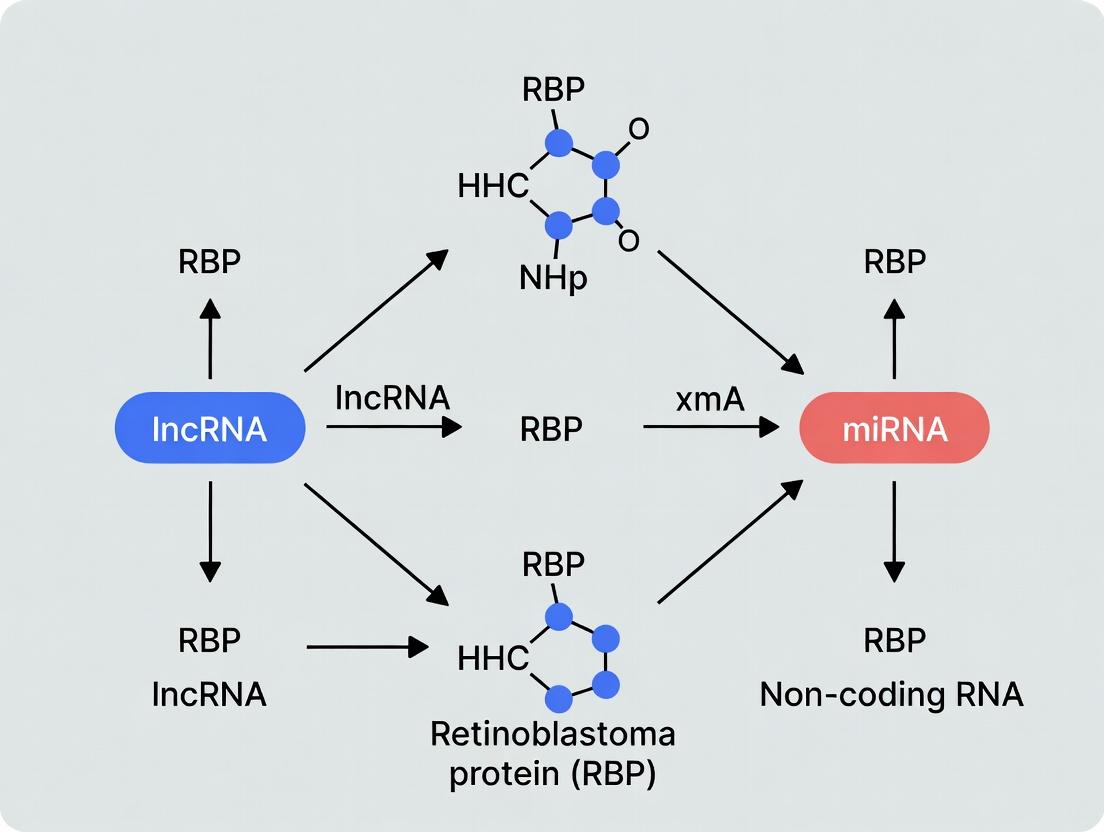
Diagram 1: Core RBP-lncRNA-miRNA interaction network.
Diagram 2: CLIP-seq experimental workflow.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Studying RBP-ncRNA Interactions
| Reagent Category | Specific Example | Function in Research |
|---|---|---|
| CLIP-grade Antibodies | Anti-AGO2, Anti-HNRNPA1 | High-specificity antibodies for immunoprecipitating endogenous RBPs in CLIP protocols. |
| Photoactivatable Nucleosides | 4-Thiouridine (4SU) | Incorporated into RNA for efficient crosslinking at 365 nm in PAR-CLIP techniques. |
| RNase Inhibitors | Recombinant RNasin / SUPERase•In | Protect RNA from degradation during cell lysis and immunoprecipitation steps. |
| Crosslinkers | Formaldehyde, UV Light (254 nm) | Form covalent bonds between interacting proteins and RNA (in vivo or in vitro). |
| Biotinylated Probes | LNA or DNA Oligonucleotides | For pull-down of specific lncRNAs or miRNAs and their associated protein complexes. |
| CRISPR Screening Libraries | dCas9-KRAB sgRNA Libraries (e.g., Brunello) | For genome-wide or focused loss-of-function screens targeting lncRNA loci. |
| NGS Library Prep Kits | SMARTer smRNA-seq Kit, CLIP-seq Kits | Optimized for constructing sequencing libraries from small RNAs or fragmented CLIP RNA. |
RNA-binding proteins (RBPs) are central arbiters of post-transcriptional gene regulation, and their interactions with non-coding RNAs (ncRNAs), particularly long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), underpin critical cellular functions and disease pathologies. This whitepaper delves into the molecular and structural principles governing RBP recognition of lncRNA and miRNA motifs, synthesizing recent high-throughput and structural data. The content is framed within the broader thesis that a mechanistic understanding of these interactions is pivotal for deciphering ncRNA function and developing RNA-targeted therapeutics.
RBPs engage with lncRNAs and miRNAs through a complex interplay of sequence, structure, and dynamics. While miRNAs present as short, often structured, single-stranded RNAs primarily recognized during their biogenesis and loading into Argonaute, lncRNAs offer expansive scaffolds for multivalent RBP interactions. Recognition is governed by modular RNA-binding domains (RBDs) such as RRMs, KH domains, and zinc fingers, which read out specific RNA signatures.
Structural Principles of Recognition
Recognition of miRNA Structures
RBPs bind miRNA precursors (pri- and pre-miRNAs) to regulate their processing by Drosha and Dicer, or to influence miRNA stability and activity.
- DGCR8/Drosha Complex (Microprocessor): DGCR8 recognizes the N6-methyladenosine (m6A) mark and the UG(U) sequence at the base of pri-miRNA stem-loop structures, facilitating accurate cropping by Drosha.
- LIN28: Binds the GGAGA motif in the terminal loop of let-7 family pre-miRNAs, recruiting a TUTase to block Dicer processing.
- Argonaute (AGO): The core RBP of the miRNA-induced silencing complex (miRISC) employs a bilobed architecture to bind the miRNA guide strand, with nucleotide specificity at positions 2-8 (seed region) critical for target mRNA recognition.
Recognition of lncRNA Structures
lncRNAs adopt complex secondary and tertiary structures, creating unique binding platforms for RBPs.
- Modular Domains and Multivalency: RBPs like HNRNPK, RBFOX2, and TDP-43 often bind short, degenerate motifs on lncRNAs. High-affinity, specific binding emerges from the cooperative binding of multiple RBDs to multiple motifs presented on a structured lncRNA scaffold (e.g., XIST, MALAT1, NEAT1).
- Shape Readout: The physical shape of the RNA groove, defined by its 3D structure, can be as important as the nucleotide sequence for RBP docking (e.g., SLBP binding to the stem-loop of histone mRNAs, a lncRNA class).
- Conditional Structures: Many lncRNAs are intrinsically disordered, folding upon binding to specific RBPs—a conformational selection mechanism.
Table 1: Quantitative Parameters of Key RBP-ncRNA Interactions
| RBP | ncRNA Target | Binding Affinity (Kd) | Recognized Motif/Structure | Key Technique Used |
|---|---|---|---|---|
| LIN28 | pre-let-7 | ~10-100 nM | GGAGA in terminal loop | EMSA, ITC |
| DGCR8 | pri-miRNA | ~30 nM (for core motif) | N6mA, UG(U) basal UG | CLIP-seq, SPR |
| HNRNPK | XIST (RepA) | Low μM (individual) | Poly-C rich sequences | RIP-seq, MST |
| TDP-43 | NEAT1 | Not well quantified | UG/GU repeats | PAR-CLIP |
| AGO2 | Mature miRNA | <1 nM (tight complex) | miRNA 5' seed & 3' end | X-ray Crystallography |
Core Experimental Methodologies
Protocol: Crosslinking and Immunoprecipitation (CLIP) and Variants
CLIP identifies genome-wide RBP-RNA interaction sites in vivo.
- In Vivo Crosslinking: Cells are irradiated with 254 nm UV-C light (150-400 mJ/cm²) to form covalent bonds between RBPs and closely bound RNAs.
- Cell Lysis and Partial RNase Digestion: Lysates are treated with limited RNase (e.g., RNase I) to fragment bound RNAs, leaving a short (~50-100 nt) protein-protected "footprint."
- Immunoprecipitation: The RBP-RNA complex is purified using specific antibodies.
- RNA Adapter Ligation and Protein Removal: 3' and 5' RNA adapters are ligated. Proteins are removed by Proteinase K digestion.
- cDNA Library Preparation & Sequencing: RNA is reverse-transcribed, amplified via PCR, and sequenced.
- Variants: PAR-CLIP uses 4-thiouridine incorporation and 365 nm crosslinking, inducing T-to-C transitions in sequencing data for precise site identification. eCLIP improves efficiency and reduces adapter contamination.
Protocol: Electrophoretic Mobility Shift Assay (EMSA)
EMSA quantifies RBP-RNA binding affinity and specificity in vitro.
- Probe Labeling: A short RNA oligonucleotide containing the putative binding site is synthesized and 5' end-labeled with [γ-³²P] ATP using T4 Polynucleotide Kinase.
- Binding Reaction: Purified recombinant RBP (serially diluted) is incubated with the labeled RNA probe (constant, low nM concentration) in a binding buffer (containing salts, carrier RNA like yeast tRNA, and RNase inhibitors) for 20-30 minutes at room temperature.
- Non-Denaturing Gel Electrophoresis: The reaction mixture is loaded onto a pre-run 4-10% polyacrylamide gel in 0.5x TBE buffer at 4°C. Protein-RNA complexes migrate slower than free RNA.
- Detection: The gel is dried and exposed to a phosphorimager screen. Fraction bound is quantified, and Kd is calculated by fitting data to a binding isotherm.
Protocol: Isothermal Titration Calorimetry (ITC)
ITC directly measures the thermodynamics of binding (Kd, ΔH, ΔS, stoichiometry) in solution.
- Sample Preparation: Highly purified RBP and RNA are dialyzed into identical buffer conditions (same pH, salt, detergent).
- Titration: The RNA solution (in the syringe) is injected in a series of small aliquots (e.g., 2-10 μL) into the RBP solution (in the cell) while stirring.
- Heat Measurement: The instrument measures the minute heat released or absorbed (μcal/sec) after each injection until the system reaches equilibrium.
- Data Analysis: The integrated heat peaks are plotted against the molar ratio. Nonlinear regression of the binding isotherm yields all thermodynamic parameters.
Visualization of Pathways and Workflows
Diagram 1: Pri-miRNA processing and RBP regulation
Diagram 2: PAR-CLIP experimental workflow
Diagram 3: RBP recognition modes for lncRNA vs. miRNA
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Studying RBP-ncRNA Interactions
| Reagent/Category | Specific Example(s) | Function & Application |
|---|---|---|
| Crosslinkers | UV-C (254 nm), 4-thiouridine + UV-A (365 nm) | Creates covalent bonds between RBPs and bound RNAs for CLIP techniques. |
| CLIP-Grade Enzymes | RNase I, Proteinase K, T4 PNK, RNase Inhibitor | Fragmentation, protein digestion, and RNA end-modification for CLIP library prep. |
| High-Affinity Antibodies | Anti-FLAG M2, Anti-HA, Anti-MYC, target-specific RBP antibodies | Immunoprecipitation of epitope-tagged or endogenous RBP-RNA complexes. |
| Next-Gen Sequencing Kits | Illumina TruSeq Small RNA Kit, SMARTer smRNA-Seq Kit | Preparation of cDNA libraries from CLIP-recovered or purified small/long RNAs. |
| Recombinant RBP Production | Baculovirus (Insect Cell), E. coli expression vectors (pET, GST) | Generates purified, active protein for in vitro assays (EMSA, ITC, crystallography). |
| Synthetic RNA Oligos | 2'-F/2'-O-Methyl modified, biotinylated, Cy5/fluorescently labeled | Probes for EMSA, pull-downs, fluorescence anisotropy, and inhibition studies. |
| In Vitro Transcription Kits | T7, SP6 RNA Polymerase kits, NTP mixes, cap analogs (for pri-miRNAs) | Produces long, structured RNA molecules (e.g., full-length lncRNA domains, pri-miRNAs). |
| Thermodynamic Analysis | MicroCal ITC instruments & consumables | Directly measures binding constants and thermodynamics in solution. |
1. Introduction This whitepaper details the functional consequences of interactions between RNA-binding proteins (RBPs) and non-coding RNAs (ncRNAs), specifically long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), within the broader thesis of RBP-ncRNA regulatory networks. These interactions are fundamental to post-transcriptional control mechanisms governing gene expression, with direct implications for cellular homeostasis and disease pathogenesis, including cancer and neurological disorders. Understanding these mechanisms is critical for identifying novel therapeutic targets in drug development.
2. Core Mechanisms of RBP-ncRNA Functional Outcomes RBPs execute their functions through defined biochemical interactions with target RNAs. The primary consequences are summarized below.
Table 1: Functional Consequences of RBP-ncRNA Interactions
| Functional Consequence | Primary Mediator | Target RNA | Key Example (RBP:RNA) | Net Effect on Gene Expression |
|---|---|---|---|---|
| mRNA Stability Control | RBP binding to AREs or other cis-elements, often in concert with miRNAs | mRNA | TTP (Tristetraprolin): ARE-containing mRNAs | Destabilization; Increased decay |
| RNA Localization & Trafficking | RBP recognition of zipcode sequences | mRNA, lncRNA | ZBP1: β-actin mRNA | Altered subcellular localization |
| miRNA/ceRNA Sponging | lncRNA or circRNA with miRNA Response Elements (MREs) | miRNA | LincRNA-ROR: miR-145 | Derepression of miRNA target mRNAs |
| Translation Regulation | RBP binding to 5'UTR or 3'UTR, influencing ribosome recruitment | mRNA | FMRP (Fragile X Mental Retardation Protein): Specific target mRNAs | Repression or Activation |
| RBP Activity Modulation | lncRNA acting as a decoy or scaffold for RBPs | RBP | MALAT1: SRSF1 (Serine/Arginine-Rich Splicing Factor 1) | Altered RBP function (e.g., splicing) |
3. Detailed Methodologies for Key Experiments Note: All protocols require appropriate controls (e.g., scramble siRNA, empty vector, IgG for IP).
3.1. Protocol: Assessing RNA Stability via Transcription Inhibition Objective: Measure the half-life of a target mRNA under conditions of RBP or lncRNA perturbation.
- Cell Transfection: Transfect cells with siRNA against the RBP/lncRNA or overexpression plasmid.
- Transcription Arrest: At 48h post-transfection, add Actinomycin D (5 µg/mL) or DRB (5,6-Dichlorobenzimidazole 1-β-D-ribofuranoside, 100 µM) to inhibit RNA polymerase II.
- Time-Course Harvest: Collect total RNA at time points (e.g., 0, 1, 2, 4, 8h) post-inhibition using TRIzol.
- Analysis: Perform RT-qPCR for the target mRNA. Normalize to a stable internal control (e.g., GAPDH or 18S rRNA). Calculate half-life (t1/2) by fitting decay data to an exponential curve.
3.2. Protocol: RNA Immunoprecipitation (RIP) and RIP-Seq Objective: Identify direct RNA targets of a specific RBP.
- Cell Lysis: Lyse cells in polysome lysis buffer (e.g., containing RNase inhibitors and protease inhibitors).
- Immunoprecipitation: Incubate lysate with antibody against the target RBP (or IgG control) conjugated to magnetic Protein A/G beads.
- Washes: Stringently wash beads with high-salt buffer to reduce non-specific binding.
- RNA Isolation: Digest proteins with Proteinase K, then extract RNA from the immunoprecipitate.
- Downstream Analysis: Analyze by RT-qPCR for candidate RNAs or prepare libraries for high-throughput sequencing (RIP-Seq).
3.3. Protocol: Cytoplasmic/Nuclear Fractionation with RNA Extraction Objective: Determine subcellular localization changes of an RNA upon RBP/lncRNA perturbation.
- Fractionation: Use a commercial kit or hypotonic lysis with NP-40 to separate cytoplasmic and nuclear fractions. Validate purity via immunoblotting (e.g., GAPDH for cytoplasm, Lamin B1 for nucleus).
- RNA Isolation: Extract RNA separately from each fraction.
- Analysis: Perform RT-qPCR for the RNA of interest in each fraction. Express as percentage of total (cytoplasmic + nuclear) signal.
4. Visualization of Pathways and Workflows
Diagram 1: RBP-ncRNA Interaction Network (Width: 760px)
Diagram 2: Experimental RIP-Seq Workflow (Width: 760px)
Diagram 3: ceRNA Sponging Mechanism (Width: 760px)
5. The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for RBP-ncRNA Functional Studies
| Reagent/Solution | Supplier Examples | Primary Function in Experiments |
|---|---|---|
| Actinomycin D | Sigma-Aldrich, Tocris | Global transcription inhibitor for RNA stability/pulse-chase assays. |
| DRB (5,6-Dichlorobenzimidazole 1-β-D-ribofuranoside) | Sigma-Aldrich, Cayman Chemical | Reversible Pol II inhibitor for RNA stability assays. |
| Proteinase K | Thermo Fisher, Qiagen | Digests proteins post-Immunoprecipitation to liberate bound RNA. |
| RNase Inhibitor (e.g., RNasin, SUPERase-In) | Promega, Thermo Fisher | Prevents RNA degradation during cell lysis and IP procedures. |
| Magnetic Protein A/G Beads | Thermo Fisher, Millipore | Solid support for antibody-based immunoprecipitation of RBPs. |
| Cytoplasmic/Nuclear Fractionation Kit | NE-PER Kit (Thermo), PARIS Kit (Thermo) | Isolates subcellular RNA compartments for localization studies. |
| 4-Thiouridine (4sU) | Sigma-Aldrich, MedChemExpress | Metabolic label for nascent RNA; enables analysis of newly transcribed RNA pools. |
| CLIP-Seq Kit (e.g., iCLIP, eCLIP) | EpiCypher, commercial protocols | Standardized reagents/protocols for high-resolution RBP-RNA crosslinking studies. |
| Locked Nucleic Acid (LNA) miRNA Inhibitors | Qiagen, Exiqon | High-affinity, nuclease-resistant inhibitors for specific miRNA functional knockdown. |
Key Biological Systems and Pathways Governed by RBP-ncRNA Networks
Within the broader thesis on RBP-ncRNA interactions, the dynamic networks formed by RNA-binding proteins (RBPs) and non-coding RNAs (ncRNAs) are established as central regulatory hubs. These networks govern core cellular processes by modulating gene expression at transcriptional, post-transcriptional, and translational levels. This technical guide details the key biological systems and signaling pathways orchestrated by these intricate RBP-ncRNA circuits, with a focus on mechanistic insights and experimental interrogation.
Core Biological Systems Regulated by RBP-ncRNA Networks
Cell Cycle Control and Proliferation
RBPs and ncRNAs form tight feedback loops to ensure precise cell cycle progression. Key nodes include the p53 pathway, cyclin-dependent kinases (CDKs), and the retinoblastoma protein (pRB) network.
Table 1: Key RBP-ncRNA Interactions in Cell Cycle Control
| RBP | ncRNA Partner | Function in Cell Cycle | Quantitative Impact (Knockdown/Overexpression) | Pathway |
|---|---|---|---|---|
| LIN28 | let-7 miRNA family | Promotes G1/S transition by repressing cell cycle inhibitors | LIN28 OE reduces let-7 levels by >80%, increasing Cyclin D1 2.5-fold. | pRB-E2F |
| HuR (ELAVL1) | p21 mRNA, Cyclin A/B mRNAs | Stabilizes mRNAs of pro-proliferative factors; destabilizes CDK inhibitors | HuR KD decreases Cyclin B1 mRNA half-life by ~60%. | G2/M Checkpoint |
| QKI | miR-20a | Modulates miRNA processing to influence E2F1 levels | QKI loss increases pre-miR-20a by 3-fold, repressing E2F1. | p53 Network |
| MSI2 | NUMB mRNA | Represses NUMB translation, promoting NOTCH signaling and proliferation | MSI2 KD reduces colony formation by 70% in assays. | NOTCH |
Apoptosis and Cellular Stress Response
These networks integrate stress signals to decide cell fate. Central pathways include mitochondrial (intrinsic) and death receptor (extrinsic) apoptosis.
Table 2: RBP-ncRNA Regulators of Apoptosis
| Regulatory Node | ncRNA Component | RBP Partner | Effect on Apoptosis | Key Target |
|---|---|---|---|---|
| Pro-survival | MALAT1 (lncRNA) | BCL2, BCL-XL mRNA (via HuR) | Inhibits apoptosis under stress | Caspase-3/9 activity reduced by 40% with MALAT1 OE. |
| Pro-apoptotic | miR-34a | p53 (transcriptional target), SIRT1 mRNA | Promotes p53 activity | miR-34a mimic increases cleaved PARP 3-fold. |
| Stress Granule | TUG1 (lncRNA) | TIA1, TIAR | Sequesters pro-apoptotic mRNAs | Oxidative stress increases RBP-TUG1 co-localization by 5x. |
| p53 Pathway | PANDAR (lncRNA) | NF-YA | Modulates p53-dependent transcription | PANDAR KD increases p53 target gene expression 2-4x. |
Immune Response and Inflammation
RBP-ncRNA networks fine-tune innate and adaptive immune signaling, notably the NF-κB and interferon (IFN) pathways.
Table 3: Immune Signaling via RBP-ncRNA Networks
| Immune Pathway | Key RBP | Interacting ncRNA | Regulatory Mechanism | Outcome |
|---|---|---|---|---|
| NF-κB Signaling | HNRNPU | NKILA (lncRNA) | lncRNA masks phosphorylation site on IκB, inhibiting NF-κB | NKILA OE reduces IL-6 secretion by 65%. |
| Type I IFN Response | Regnase-1 | IL6 mRNA, TNFA mRNA | Endonuclease degrades inflammatory mRNAs | Regnase-1 KO increases serum IL-6 >10x. |
| NLRP3 Inflammasome | DDX3X | NEAT1 (lncRNA) | Facilitates NLRP3 transcription and inflammasome assembly | DDX3X inhibitor decreases IL-1β release by 80%. |
| T Cell Differentiation | ROQUIN | ICOS mRNA, miR-101 | Degrades mRNAs of activating receptors; modulates miRNA. | ROQUIN mutation leads to autoimmunity. |
Cellular Differentiation and Development
These networks provide cell fate-determining post-transcriptional programs, essential in stemness and tissue development.
Table 4: RBP-ncRNA in Differentiation
| System | RBP | ncRNA | Target/Pathway | Differentiation Role |
|---|---|---|---|---|
| Neurogenesis | MUSASHI (MSI1) | mRNA of Numb, Notch | Represses translation of differentiation promoters | MSI1 loss triggers premature differentiation. |
| Myogenesis | LIN28 | let-7 miRNA | Blocks let-7 biogenesis, maintaining progenitor state | LIN28A OE enhances muscle regeneration 2x. |
| Hematopoiesis | IGF2BP1-3 | H19, XIST (lncRNAs), MYC mRNA | Stabilizes MYC mRNA; interacts with lncRNAs | IGF2BP1 KD reduces CFU assays by 90%. |
| Adipogenesis | HuR | miR-let-7d, PPARγ mRNA | Competes with let-7d for PPARγ mRNA binding | HuR inhibition reduces lipid accumulation by 60%. |
Detailed Experimental Protocols
Protocol: Crosslinking and Immunoprecipitation (CLIP) for Mapping RBP-ncRNA Interactions
Purpose: To identify genome-wide binding sites of an RBP on its RNA targets in vivo. Key Reagents: UV crosslinker (254 nm), Protein A/G magnetic beads, RNAse T1, Phosphatase, PNK enzyme, 3' RNA adaptor, 5' RNA adaptor, High-fidelity reverse transcriptase, Proteinase K.
- In Vivo Crosslinking: Culture cells (~1x10^7 per condition). Irradiate with 254 nm UV light (400 mJ/cm²) to covalently link RBPs to bound RNA.
- Cell Lysis: Lyse cells in stringent RIPA buffer with RNAse inhibitors.
- Partial RNA Digestion: Treat lysate with RNAse T1 (0.01 U/µL) to digest unbound RNA, leaving ~20-60 nt protected fragments.
- Immunoprecipitation: Pre-clear lysate. Incubate with antibody against target RBP (or control IgG) bound to magnetic beads for 2-4h at 4°C.
- 3' Dephosphorylation & Ligation: Wash beads. Dephosphorylate RNA 3' ends with phosphatase. Ligate a pre-adenylated 3' DNA adaptor using T4 RNA Ligase 1 (truncated).
- Radioactive 5' Phosphorylation (Optional): For visualization, use PNK with γ-³²P-ATP to label 5' ends.
- Proteinase K Digestion & RNA Recovery: Elute RBP-RNA complexes in SDS buffer. Digest protein with Proteinase K. Phenol-chloroform extract RNA.
- 5' Adaptor Ligation & Reverse Transcription: Ligate 5' RNA adaptor. Reverse transcribe using SuperScript IV.
- Library Prep & Sequencing: Amplify cDNA by PCR. Size-select and sequence on an Illumina platform.
- Bioinformatic Analysis: Map reads to genome, call peaks (e.g., using CLIPper, PEAKachu), and identify binding motifs.
Protocol: RNA Immunoprecipitation (RIP) and RIP-Seq
Purpose: To identify RNAs associated with a specific RBP under native (non-crosslinked) conditions. Key Reagents: RIP buffer (with Mg2+, RNAse inhibitors), Magnetic beads (Protein A/G), DNase I, RNA clean-up kit, NGS library prep kit.
- Cell Lysis: Lyse cells in mild RIP lysis buffer to preserve native interactions.
- Immunoprecipitation: Incubate pre-cleared lysate with specific antibody-coated beads (or control) overnight at 4°C.
- Washing & Elution: Wash beads 5x with RIP wash buffer. Elute bound RNA-protein complexes using buffer containing SDS and Proteinase K.
- RNA Extraction: Purify RNA using phenol-chloroform or a spin column kit. Treat with DNase I.
- Analysis:
- qRT-PCR: For candidate RNAs.
- RIP-Seq: Prepare RNA-seq library from immunoprecipitated and total input RNA. Sequence. Enriched RNAs are identified by comparing IP vs. input signals (e.g., using DESeq2).
Protocol: Functional Validation Using CRISPR/Cas9 and Reporter Assays
Purpose: To validate the regulatory consequence of a specific RBP-ncRNA interaction on a target pathway. Key Reagents: sgRNAs targeting RBP or ncRNA locus, Cas9 expression plasmid, Dual-Luciferase Reporter Assay System, Target gene 3'UTR luciferase construct.
- Generate Knockout Cell Line: Co-transfect cells with Cas9 and sgRNA plasmids. Select with puromycin for 72h. Single-cell clone and validate KO by western blot (RBP) or PCR (lncRNA).
- Reporter Assay:
- Clone the putative target 3'UTR (or ncRNA sequence) downstream of a Renilla luciferase gene in a psiCHECK2 vector.
- Co-transfect WT and KO cells with the reporter plasmid and a Firefly luciferase control plasmid.
- After 48h, lyse cells and measure luminescence using a Dual-Luciferase Assay kit.
- Calculate the ratio of Renilla (experimental) to Firefly (control). A change in ratio in KO vs. WT confirms functional regulation.
Visualization of Pathways and Workflows
Diagram 1: Cell Cycle Regulation by RBP-ncRNA Networks
Diagram 2: CLIP-Seq Workflow for Mapping RBP Binding Sites
Diagram 3: NF-κB Pathway Modulation by lncRNA NKILA
The Scientist's Toolkit: Essential Research Reagents
Table 5: Key Research Reagent Solutions
| Reagent Category | Specific Example(s) | Function in RBP-ncRNA Research |
|---|---|---|
| Crosslinkers | UV-C (254 nm) light source; Formaldehyde | UV creates covalent protein-RNA bonds for CLIP. Formaldehyde captures indirect/complex interactions. |
| Immunoprecipitation Beads | Protein A, Protein G, or A/G Magnetic Beads | Solid support for capturing antibody-RBP complexes during RIP or CLIP. Magnetic beads facilitate washing. |
| Ribonucleases | RNase T1, RNase A, RNase I | Used in CLIP to digest unprotected RNA, leaving protein-bound footprints. Different specificities inform on binding mode. |
| Adaptor Ligases | T4 RNA Ligase 1 (truncated K227Q), T4 RNA Ligase 2 | Ligate DNA/RNA adaptors to RNA ends for sequencing library construction. Truncated Ligase 1 prefers pre-adenylated 3' adaptors. |
| High-Fidelity Enzymes | SuperScript IV RT, Phusion HF DNA Polymerase | Critical for accurate reverse transcription and PCR amplification of low-yield, crosslinked RNA fragments. |
| CLIP-Seq Kits | iCLIP2, eCLIP Commercial Kits | Standardized reagent sets that improve efficiency and reproducibility of library preparation for high-throughput sequencing. |
| CRISPR/Cas9 Components | Cas9 Nuclease, sgRNAs, HDR templates | For generating knockout cell lines of specific RBPs or ncRNA loci to study loss-of-function phenotypes. |
| Dual-Luciferase Reporter Systems | psiCHECK2, pmirGLO Vectors | Quantify the regulatory impact of an RBP-ncRNA interaction on a specific mRNA target's stability or translation. |
| RNA-Binding Protein Arrays | Protein array membranes with spotted RBPs | High-throughput screening for identifying RBP partners of a labeled ncRNA probe. |
| In Situ Hybridization (ISH) Probes | ViewRNA, RNAscope Probe Sets | Visualize the subcellular localization of ncRNAs in fixed cells/tissues, often combined with IF for RBPs. |
Evolutionary Perspectives on the Conservation of RBP-ncRNA Interactions
The intricate network of interactions between RNA-binding proteins (RBPs) and non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), forms a critical regulatory layer in cellular biology. From an evolutionary standpoint, the conservation of these interactions across species provides a powerful lens through which to understand functional importance, identify essential regulatory nodes, and prioritize targets for therapeutic intervention in conservation biology and disease. This whitepaper, framed within the broader thesis of RBP-ncRNA interplay, examines how evolutionary principles guide the conservation of these interactions, offering a technical guide for researchers and drug development professionals.
Evolutionary Principles and Conservation Metrics
The conservation of an RBP-ncRNA interaction implies selective pressure to maintain the interaction interface—the specific sequence or structural motifs in the RNA and the corresponding binding domains in the protein. Key evolutionary metrics are used to assess this conservation.
Table 1: Key Evolutionary Metrics for Assessing RBP-ncRNA Interaction Conservation
| Metric | Description | Application in RBP-ncRNA Context |
|---|---|---|
| Phylogenetic Footprinting | Identification of conserved sequence blocks across species. | Identifies conserved RNA motifs potentially serving as RBP binding sites. |
| Sequence Identity (%) | Percentage of identical nucleotides (or amino acids) between orthologous sequences. | High identity in binding regions suggests functional constraint. |
| Synonymous vs. Non-synonymous Substitution Rate (dN/dS) | Ratio of nucleotide changes altering amino acids (non-synonymous) to silent changes (synonymous). | dN/dS < 1 in an RBP's RNA-binding domain or ncRNA's protein-binding motif indicates purifying selection. |
| Conservation Score (e.g., PhastCons, Gerp++) | Genomic evolutionary rate profiling scores quantifying nucleotide-level constraint. | High scores in specific ncRNA regions predict functionally important, and likely protein-bound, elements. |
| Co-evolution Analysis | Detection of correlated evolutionary changes between interacting partners. | Compensatory mutations in RBP and its ncRNA partner signal maintained interaction. |
Experimental Protocols for Studying Conservation
Protocol: Cross-Species CLIP-Seq (Crosslinking and Immunoprecipitation)
Objective: To experimentally map RBP binding sites on ncRNAs across different species and assess binding site conservation. Detailed Methodology:
- Cell/Organism Selection: Choose phylogenetically spaced species (e.g., human, mouse, chicken, zebrafish).
- In Vivo Crosslinking: Expose cells/tissues to UV-C light (254 nm) to covalently crosslink RBPs to bound RNA.
- Cell Lysis and Immunoprecipitation: Lyse cells and immunoprecipitate the RBP of interest using a specific antibody.
- RNA Processing: Treat with RNase to partially digest unbound RNA, leaving protected protein-bound fragments. Dephosphorylate and ligate a 3' adapter.
- Isolation and cDNA Synthesis: Isolate RBP-RNA complexes by SDS-PAGE, then release and purify RNA fragments. Ligate a 5' adapter, reverse transcribe to cDNA, and PCR amplify.
- High-Throughput Sequencing: Sequence libraries and map reads to respective reference genomes.
- Comparative Bioinformatics Analysis: Identify significant binding peaks. Use liftOver tools and multiple sequence alignments to identify orthologous genomic regions in other species. Test for enrichment of binding in conserved genomic blocks.
Protocol: Electrophoretic Mobility Shift Assay (EMSA) with Orthologous RNAs
Objective: To quantitatively compare the binding affinity of an RBP for its cognate ncRNA target across different species. Detailed Methodology:
- RNA Probe Preparation: In vitro transcribe and purify the ncRNA region of interest from multiple species (e.g., human XIST RepA, mouse Xist RepA). Label with [γ-32P] ATP or a fluorescent dye.
- Protein Purification: Express and purify the recombinant RBP (e.g., PRC2 complex) from a heterologous system.
- Binding Reactions: Incubate a fixed amount of labeled RNA probe with increasing concentrations of the RBP (0-500 nM) in binding buffer (e.g., containing heparin, tRNA, DTT) for 20-30 minutes at room temperature.
- Non-Denaturing Gel Electrophoresis: Load reactions onto a pre-run 4-10% polyacrylamide gel in 0.5x TBE buffer. Run at 4°C to maintain complexes.
- Detection and Analysis: Visualize shifted complexes (RBP-bound RNA) and free probe via phosphorimaging or fluorescence. Calculate apparent dissociation constant (Kd) for each orthologous RNA using non-linear regression analysis of binding curves.
- Conservation Interpretation: A similar low Kd across species indicates a conserved, high-affinity interaction. A significantly increased Kd in a distant species suggests divergence or loss of function.
Visualization of Key Concepts and Workflows
Diagram Title: Evolutionary Conservation Analysis Pipeline
Diagram Title: Co-evolution and Divergence of RBP-ncRNA Pairs
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Studying RBP-ncRNA Conservation
| Item | Function in Conservation Studies |
|---|---|
| Species-Specific RBP Antibodies | For immunoprecipitation in cross-species CLIP-seq experiments. Validated for the target species is critical. |
| Orthologous ncRNA Expression Vectors | Plasmids to express the ncRNA from different species in a common cellular background for functional comparison. |
| Recombinant Orthologous RBPs | Purified proteins from multiple species for in vitro binding affinity assays (EMSA, SPR). |
| CRISPR/Cas9 Reagents for Multiple Species | Guide RNAs and Cas9 constructs to knockout or mutate the RBP or ncRNA locus in various model organisms. |
| Phylogenetic Analysis Software (e.g., PhyloP, PHAST) | Computes conservation scores from multiple sequence alignments to identify constrained regions. |
| Crosslinking Equipment (UV Lamp, 254 nm) | Standardized equipment for in vivo or in vitro crosslinking to capture transient RBP-ncRNA interactions. |
| Nucleotide Analogs (4-Thiouridine, 6-Thioguanosine) | For PAR-CLIP protocols, introduces specific mutation signatures for precise binding site identification. |
| Stable Cell Lines Expressing Tagged RBPs | Enables consistent pull-down across experiments; tagging must not disrupt the native interaction. |
Implications for Conservation and Drug Development
Evolutionarily conserved RBP-ncRNA interactions represent fundamental regulatory modules. In conservation biology, their disruption can serve as biomarkers for environmental stress. In drug development, they present high-value, genetically validated targets. Small molecules or oligonucleotides (ASOs, siRNAs) designed to disrupt a conserved, disease-driving interaction offer high potential for efficacy with a potentially wider therapeutic window, as the targeted interface is likely under strong functional constraint. Conversely, species-specific interactions may explain side effects or lack of translation from animal models, highlighting the necessity of evolutionary analysis in preclinical research.
From Detection to Discovery: Cutting-Edge Methods to Map and Analyze RBP-ncRNA Interactomes
Within the study of RNA-binding protein (RBP) interactions with non-coding RNAs (lncRNAs, miRNAs), precise mapping of binding sites is paramount. This technical guide details the core methodologies—RIP-seq and its higher-resolution CLIP-seq variants—that form the gold standard for in vivo RBP-RNA interaction analysis. These techniques are foundational for elucidating post-transcriptional regulatory networks in development, disease, and drug discovery.
Core Methodologies & Quantitative Comparison
RIP-seq (RNA Immunoprecipitation followed by sequencing) identifies RNAs associated with a specific RBP but does not provide nucleotide-resolution binding sites. CLIP-seq (Crosslinking and Immunoprecipitation) and its derivatives overcome this by incorporating UV crosslinking to covalently link RBPs to their bound RNAs, allowing for precise mapping after rigorous purification.
Quantitative Comparison Table
Table 1: Comparative Overview of RIP-seq and CLIP-seq Variants
| Feature | RIP-seq | HITS-CLIP | PAR-CLIP | iCLIP |
|---|---|---|---|---|
| Crosslinking | None (native) | UV-C (254 nm) | UV-A (365 nm) + 4-Thiouridine | UV-C (254 nm) |
| Resolution | Transcript-level | ~30-60 nt | ~20-30 nt | Single-nucleotide |
| Key Mutational Signal | N/A | Crosslink-induced deletions | T-to-C transitions | cDNA truncations |
| Primary Output | Enriched transcripts | Binding regions | Binding sites with mutations | Precise crosslink sites |
| Typical SNR* | Low | Moderate | High | High |
| Application in ncRNA | Identification of bound lncRNAs/miRNAs | Mapping interactions on long RNAs | Ideal for miRNA binding sites | Protein-RNA interface studies |
*SNR: Signal-to-Noise Ratio
Detailed Experimental Protocols
Protocol 1: Core CLIP-seq Workflow
- In Vivo Crosslinking: Cells are irradiated with UV light (254 nm for HITS-CLIP/iCLIP, 365 nm for PAR-CLIP-treated cells).
- Cell Lysis: Use stringent lysis buffers (e.g., containing RIPA, RNase inhibitors, protease inhibitors).
- Partial RNase Digestion: Treat lysate with limited RNase to fragment bound RNA, leaving ~20-60 nt footprints.
- Immunoprecipitation (IP): Use antibody-coated beads specific to the RBP of interest.
- RNA Linker Ligation: For iCLIP and HITS-CLIP, a 3' RNA adapter is ligated after dephosphorylation.
- Protein Removal & RNA Isolation: Treat with Proteinase K, purify RNA via phenol-chloroform extraction.
- cDNA Library Prep: Reverse transcribe, ligate 5' adapter, PCR amplify, and sequence.
Protocol 2: PAR-CLIP-Specific Steps
- Prior to crosslinking, cells are grown in medium supplemented with 4-thiouridine (4SU).
- Crosslink with UV-A at 365 nm, which induces efficient crosslinking between the RBP and 4SU.
- During reverse transcription, crosslinked 4SU residues cause T-to-C transitions in the cDNA, providing a diagnostic mutation signal.
Protocol 3: iCLIP-Specific Steps
- After IP, RNAs are dephosphorylated and a 3' ssDNA linker is ligated directly to the RNA.
- During reverse transcription, the enzyme frequently stops at the crosslink site, producing truncated cDNAs.
- A specialized circularization and re-linearization step allows for amplification of these truncated cDNAs, pinpointing the crosslink nucleotide.
Visualizing Workflows and Concepts
Diagram 1: General CLIP-seq Experimental Workflow
Diagram 2: Differentiating CLIP-seq Variants
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CLIP-seq Experiments
| Item | Function | Example/Note |
|---|---|---|
| UV Crosslinker | Induces covalent bonds between RBPs and RNA. | UV-C (254 nm) standard; PAR-CLIP requires UV-A (365 nm). |
| 4-Thiouridine (4SU) | Photosensitive nucleoside for efficient PAR-CLIP crosslinking. | Added to cell culture medium prior to UV-A crosslinking. |
| RNase Inhibitors | Prevent exogenous RNA degradation during lysis and IP. | Recombinant RNase inhibitors (e.g., RNasin). |
| Sequence-Specific Antibodies | Immunoprecipitate the target RBP. | High specificity is critical; monoclonal antibodies preferred. |
| Magnetic Protein A/G Beads | Solid support for antibody-mediated capture of RBP-RNA complexes. | Enable efficient washing steps. |
| Partial RNase (e.g., RNase I) | Fragments unbound RNA to leave protected footprints. | Concentration must be titrated for optimal fragmentation. |
| T4 Polynucleotide Kinase (PNK) | For 5' phosphorylation/dephosphorylation in adapter ligation steps. | Essential for iCLIP workflow. |
| Truncated RNA Ligase 2 (T4 Rnl2) | Ligates pre-adenylated 3' adapters to RNA with high efficiency. | Minimizes adapter dimer formation. |
| Reverse Transcriptase | Generates cDNA from immunoprecipitated RNA fragments. | Must have high processivity and read-through capability. |
| USER Enzyme | Used in iCLIP to digest the cDNA and allow circularization. | Specific to uracil-containing DNA. |
Applications in lncRNA and miRNA Research
These techniques are pivotal for decoding RBP-ncRNA interactions. RIP-seq can survey which lncRNAs or pre-miRNAs are bound by an RBP. HITS-CLIP maps binding regions on long lncRNAs. PAR-CLIP's high resolution is excellent for defining miRNA binding sites on targets or RBPs that interact with miRNAs. iCLIP's single-nucleotide precision can reveal how an RBP's structural domains contact specific bases in an miRNA seed region or lncRNA structural motif. This data is indispensable for constructing interaction networks relevant to diseases like cancer and neurodegeneration, offering targets for therapeutic intervention.
This whitepaper details three emerging, high-resolution technologies—RAP-MS, TRIBE, and CRISPR-based screening—that are revolutionizing the study of functional interactions between RNA-binding proteins (RBPs) and non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs). Within the broader thesis of understanding the RBP-ncRNA interactome, these tools address the critical need to move beyond merely cataloguing interactions to defining their in vivo functional consequences and regulatory networks.
RAP-MS: RNA Antisense Purification with Mass Spectrometry
RAP-MS is a method for the unbiased identification of proteins that interact directly with a specific RNA molecule in vivo, without requiring prior knowledge of the protein's RNA-binding capability.
Detailed Protocol
- Design and Transfection: Design approximately 120-mer biotinylated antisense DNA oligonucleotides tiled across the target lncRNA. Co-transfect these oligonucleotides into cells alongside a psoralen derivative (e.g., AMT).
- In Vivo Crosslinking: Upon irradiation with 365 nm UVA light, psoralen crosslinks the biotinylated DNA probes to the target RNA.
- Cell Lysis and Capture: Lyse cells under denaturing conditions. Capture the RNA-DNA hybrid complex using streptavidin-coated magnetic beads.
- Stringent Washing: Wash beads stringently with denaturing buffers to eliminate non-specific protein associations.
- Elution and Digestion: Elute the protein components via RNase digestion or reversal of crosslinks. Perform on-bead tryptic digestion.
- Mass Spectrometry & Analysis: Analyze peptides by LC-MS/MS. Compare protein abundance against control oligonucleotide captures to identify specific interactors.
Key Applications
- Mapping the protein interactome of low-abundance nuclear lncRNAs (e.g., Xist, MALAT1).
- Identifying novel RBPs that bind to viral or circular RNAs.
- Characterizing changes in RNP composition under different cellular stresses.
TRIBE: Targets of RNA-Binding Proteins Identified by Editing
TRIBE (and its derivative, STAMP) employs a catalytic-dead RNA editor (dADAR) fused to an RBP of interest to mark its endogenous RNA binding sites by introducing detectable mutations (A-to-I edits) in the target transcripts.
Detailed Protocol
- Construct Generation: Create an expression vector for the RBP of interest fused to the catalytic domain of dADAR(E→Q).
- Cell Line Generation: Stably express the RBP-dADAR fusion in your model cell line or organism.
- RNA Isolation and Sequencing: Isolate total RNA and perform high-depth RNA-seq.
- Variant Calling: Use specialized alignment and variant-calling pipelines (e.g., GIREMI, REDtoolbox) to identify A-to-G mismatches (the hallmark of A-to-I editing) in the sequencing data.
- Target Identification: Statistically compare editing sites in the experimental condition to control cells expressing dADAR alone. Clusters of editing sites define true RBP binding sites.
Key Applications
- Identifying in vivo mRNA targets of RBPs at single-nucleotide resolution.
- Profiling RBP activity in specific subcellular compartments (e.g., by adding localization signals to the fusion).
- Studying dynamic changes in RBP binding across developmental stages or disease states.
CRISPR-Based Screening for Functional Interactors
CRISPR knockout (KO) or interference (CRISPRi) screens are used to perform loss-of-function genetic screens to identify genes that are functionally relevant to an RBP-ncRNA regulatory axis.
Detailed Protocol (Pooled CRISPRi Screen for RBP Genetic Interactors)
- Reporter Design: Create a fluorescent (e.g., GFP) or selectable (e.g., puromycin resistance) reporter gene under the control of a regulatory element known to be influenced by the RBP-lncRNA complex.
- Library Transduction: Transduce a cell line harboring the reporter and expressing dCas9-KRAB with a genome-wide sgRNA library (e.g., Brunello library).
- Selection & Sorting: Apply relevant selection (e.g., puromycin) and then FACS-sort cells based on reporter signal (e.g., high GFP vs. low GFP).
- Sequencing & Analysis: Recover genomic DNA from sorted populations, amplify the sgRNA barcodes via PCR, and sequence them. Use MAGeCK or similar algorithms to identify sgRNAs enriched/depleted in the phenotype of interest.
Key Applications
- Identifying genetic suppressors or enhancers of an RBP-ncRNA-mediated phenotype (e.g., cell proliferation, drug resistance).
- Mapping synthetic lethal interactions with RBP loss in cancer models.
- Validating hits from RAP-MS/TRIBE by testing their functional necessity.
Table 1: Comparative Analysis of Emerging RBP-ncRNA Interaction Tools
| Feature | RAP-MS | TRIBE/STAMP | CRISPR-based Screening |
|---|---|---|---|
| Primary Output | Proteins bound to a specific RNA | RNA targets of a specific RBP | Functional genes in an RBP/ncRNA pathway |
| Resolution | RNA-level (identifies protein binders) | Single-nucleotide (identifies binding sites) | Gene-level (identifies functional players) |
| Context | In vivo interaction, requires crosslinking | In vivo activity | Functional consequence of loss-of-function |
| Throughput | Medium (one RNA at a time) | Medium (one RBP at a time) | High (genome-wide) |
| Key Metric | Spectral counts or fold-change vs. control | Editing rate (% of reads with A-to-G change) | sgRNA log2 fold-change & p-value |
| Typical Validation | Western blot, CLIP | Independent RNA-seq, RIP-qPCR | Individual sgRNA knockout, rescue assays |
Table 2: Example Quantitative Outcomes from Representative Studies
| Tool | Target | Key Quantitative Finding | Reference Context |
|---|---|---|---|
| RAP-MS | Human Xist lncRNA | Identified >80 proteins enriched >2-fold over vector control, including 10 novel interactors. | (Minnoye et al., 2024) |
| TRIBE | Drosophila RBP Hrp48 | Detected a median editing rate of 0.3% at high-confidence binding sites vs. 0.01% background. | (Zhang et al., 2023) |
| CRISPRi Screen | LIN28A RBP | Identified 47 genes whose knockdown suppressed let-7 miRNA inhibition (FDR < 0.05). | (Liang et al., 2024) |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Implementing Featured Technologies
| Reagent / Material | Function & Importance | Example Product/Catalog |
|---|---|---|
| Psoralen (AMT) | In vivo crosslinker for RAP-MS; enables covalent bond between biotinylated DNA probe and target RNA. | 4'-Aminomethyltrioxsalen hydrochloride (Sigma A4330) |
| Streptavidin Magnetic Beads (MyOne C1) | Capture matrix for RAP-MS; high binding capacity and low non-specific binding under denaturing conditions. | Dynabeads MyOne Streptavidin C1 (Thermo 65001) |
| Catalytic-dead ADAR (dADAR) | Engineered enzyme core for TRIBE; performs binding-dependent, catalytic-independent RNA editing. | pCAG-dADAR1(E→Q)-FLAG plasmid (Addgene #102786) |
| Genome-wide sgRNA Library | Pooled guide RNA collection for CRISPR screens; enables systematic loss-of-function. | Human Brunello CRISPRko Library (Addgene #73178) |
| dCas9-KRAB Fusion Protein | Transcriptional repressor for CRISPRi screens; enables reversible gene silencing without double-strand breaks. | lenti-dCas9-KRAB-blast plasmid (Addgene #89567) |
| High-Sensitivity MS-Grade Trypsin | Protease for on-bead digestion in RAP-MS; generates peptides for LC-MS/MS analysis. | Trypsin Platinum, Mass Spec Grade (Promega VA9000) |
| UMI RNA-seq Kit | Library prep for TRIBE; incorporates Unique Molecular Identifiers to accurately quantify editing rates. | SMARTer Stranded Total RNA-Seq Kit v3 (Takara Bio 634485) |
Visualized Workflows and Pathways
RAP-MS Experimental Workflow Diagram
TRIBE/STAMP Mechanism Diagram
Pooled CRISPR Screening Workflow
Integrated Tool Strategy for RBP-ncRNA Research
Within the broader thesis on RBP interactions with non-coding RNAs, the precise mapping of protein binding sites on lncRNAs and miRNAs is a foundational challenge. In silico prediction algorithms have become indispensable for formulating hypotheses, guiding experimental design, and interpreting high-throughput data. This guide details the core algorithms, their underlying methodologies, and practical protocols for their application in research and therapeutic development.
Core Algorithmic Approaches and Quantitative Performance
Prediction algorithms can be categorized by their methodological approach. Performance is typically measured using metrics like Area Under the Curve (AUC), accuracy (Acc), precision (Prec), and recall, often benchmarked on datasets from CLIP-seq (e.g., eCLIP, PAR-CLIP) experiments.
Table 1: Comparison of Major RBP Binding Site Prediction Algorithms
| Algorithm Name | Core Methodology | Input Features | Reported Performance (Avg.) | Best For |
|---|---|---|---|---|
| DeepBind | Convolutional Neural Networks (CNN) | RNA sequence (k-mers) | AUC: 0.89-0.92 | Sequence-specific motif discovery |
| GraphProt | Support Vector Machines (SVM) with sequence/structure profiles | Sequence, predicted structure propensity | AUC: 0.85-0.90 | Modeling structure preference |
| pysster | CNN with model interpretation | Sequence, secondary structure one-hot encoding | AUC: 0.90-0.93 | Interpretable motif and structure logos |
| PRIdictor | Random Forest | k-mer sequence, RNAfold free energy | Acc: ~84%, Prec: ~0.81 | Incorporating thermodynamic stability |
| DeepCLIP | CNN/BiLSTM hybrid | Nucleotide sequence | AUC: >0.90 on eCLIP data | Generalizable models from CLIP data |
| Piano | Ensemble of multiple SVM models | Sequence, structure, conservation | AUC: 0.87-0.91 | Multi-feature, high-confidence predictions |
Experimental Protocols for Validation
Protocol: In Vitro Validation using RNA Electromobility Shift Assay (REMSA)
Purpose: To biochemically validate predicted RBP-RNA interactions. Reagents:
- Purified RBP: Recombinant protein (e.g., His-tag purified).
- RNA Probes: 20-40 nt RNA sequences containing predicted binding sites and mutant controls (Cy5-labeled at 5' end).
- Binding Buffer: 10 mM HEPES (pH 7.3), 20 mM KCl, 1 mM MgCl2, 1 mM DTT, 5% glycerol, 0.1 µg/µL yeast tRNA, 10 U/mL RNase inhibitor.
- Non-denaturing Polyacrylamide Gel: 6-8% gel in 0.5x TBE buffer. Procedure:
- Binding Reaction: Incubate 10-50 fmol of labeled RNA with increasing concentrations (0-500 nM) of purified RBP in 20 µL binding buffer for 30 min at room temperature.
- Electrophoresis: Load reactions onto pre-run 6-8% native PAGE gel in 0.5x TBE at 4°C. Run at 100 V for 60-90 min.
- Detection: Visualize gel using a fluorescence imager (Cy5 channel). A mobility shift (retarded band) indicates complex formation.
- Competition Assay: Repeat with 100x molar excess of unlabeled wild-type or mutant RNA to confirm specificity.
Protocol: In Vivo Validation using Crosslinking and Immunoprecipitation (CLIP) qPCR
Purpose: To confirm in vivo binding at predicted sites within a cellular context. Reagents:
- Cells: Relevant cell line expressing the RBP of interest.
- Crosslinker: UV-C (254 nm) light source for PAR-CLIP (or UV-A for iCLIP).
- Lysis/IP Buffer: 50 mM HEPES (pH 7.5), 150 mM KCl, 2 mM EDTA, 1% NP-40, 0.5% Sodium deoxycholate, protease/RNase inhibitors.
- Antibodies: Specific antibody against the RBP and isotype control IgG.
- Proteinase K: For digesting protein post-IP.
- TRIzol Reagent: For RNA extraction.
- Primers: qPCR primers specific for the lncRNA/miRNA region containing the predicted site. Procedure:
- Crosslinking: Wash cells with PBS and irradiate once with 254 nm UV light (400 mJ/cm²). Harvest cells.
- Cell Lysis and Immunoprecipitation: Lyse cells in IP buffer. Pre-clear lysate, then incubate with antibody-coated magnetic beads overnight at 4°C.
- Washing and Elution: Wash beads stringently. Treat with Proteinase K to reverse crosslinks and release RNA.
- RNA Extraction: Extract RNA using TRIzol.
- cDNA Synthesis and qPCR: Reverse transcribe and perform quantitative PCR using site-specific primers. Enrichment is calculated relative to input and IgG control samples (ΔΔCt method).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents and Kits for RBP-RNA Interaction Studies
| Reagent/KIT Name | Supplier Examples | Function in RBP Binding Site Research |
|---|---|---|
| Magna RIP Kit | MilliporeSigma | Optimized buffers and beads for RNA immunoprecipitation (RIP) from cells. |
| Pierce Anti-HA Magnetic Beads | Thermo Fisher | For immunoprecipitation of HA-tagged RBPs expressed in cells. |
| Dynabeads His-Tag Isolation & Pulldown | Thermo Fisher | For purifying and pulling down recombinant His-tagged RBPs for in vitro assays. |
| Silencer siRNA Kit | Thermo Fisher | For knocking down specific RBPs to study functional consequences on target ncRNAs. |
| MAXIscript T7 Transcription Kit | Thermo Fisher | For synthesizing labeled or unlabeled RNA probes for REMSA. |
| NEBNext Multiplex Small RNA Library Prep Kit | New England Biolabs | For preparing libraries from CLIP-derived RNA for high-throughput sequencing validation. |
| RNase Inhibitor, Murine | NEB, Takara | Essential for preventing RNA degradation in all biochemical assays. |
| CircLigase ssDNA Ligase | Lucigen | Critical for circularizing RNA in some CLIP library protocols (e.g., iCLIP). |
Visualized Workflows and Pathways
In Silico Prediction Workflow
CLIP to Algorithm Training Pipeline
Functional Outcomes of RBP-ncRNA Binding
This whitepaper details integrative multi-omics methodologies within the specific research framework of elucidating RNA-binding protein (RBP) interactions with non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs). Dysregulation of these interactions is implicated in cancer, neurodegenerative diseases, and metabolic disorders. A singular omics layer provides an incomplete picture; transcriptomics identifies ncRNA expression, but not their protein partners or functional outcomes. Integrative analysis is therefore critical to map the complete regulatory network from RNA expression (Transcriptomics) to protein binding and abundance (Proteomics), enabled by computational integration (Bioinformatics).
Core Multi-Omics Workflow for RBP-ncRNA Research
A synergistic experimental and computational pipeline is required to move from correlation to causation.
Diagram 1: Integrative Multi-Omics Workflow for RBP-ncRNA Studies
Detailed Experimental Protocols
Transcriptomics: Capturing ncRNA Expression Profiles
- Method: Total RNA Sequencing (RNA-seq) coupled with small RNA-seq.
- Protocol Outline:
- Sample Preparation: Extract total RNA from control and perturbed (e.g., RBP knockdown) cells using a column-based kit with DNase I treatment. Assess integrity (RIN > 8.5).
- Library Prep:
- For lncRNAs/mRNAs: Use a ribosomal RNA depletion kit, followed by fragmentation, cDNA synthesis, and adapter ligation.
- For miRNAs: Size-select small RNAs (<200 nt), ligate 3' and 5' adapters, reverse transcribe, and amplify.
- Sequencing: Pool libraries and sequence on an Illumina platform (e.g., NovaSeq). Target >40 million paired-end 150bp reads for RNA-seq and >10 million single-end 75bp reads for small RNA-seq.
- Primary Analysis: Align reads to the reference genome (e.g., GRCh38) using
STAR(RNA-seq) orBowtie2(small RNA-seq). Quantify gene expression (lncRNA, mRNA) withfeatureCountsand miRNA counts withmiRDeep2.
Proteomics: Identifying RBP Interactors and Global Protein Changes
- Method A (RBP-centric): RNA Immunoprecipitation-Mass Spectrometry (RIP-MS).
- Cell Lysis: Lyse cells in polysome lysis buffer (supplemented with RNase inhibitors and protease inhibitors).
- Immunoprecipitation (IP): Incubate lysate with antibody against target RBP or control IgG, conjugated to magnetic beads. Wash stringently.
- Elution & Digestion: Elute bound ribonucleoprotein complexes. Digest proteins with trypsin.
- LC-MS/MS: Analyze peptides by liquid chromatography-tandem mass spectrometry (e.g., Q Exactive HF).
- Method B (Global): Tandem Mass Tag (TMT) Proteomics.
- Protein Extraction & Digestion: Lyse cells in SDS buffer, reduce, alkylate, and digest with trypsin.
- TMT Labeling: Label peptides from different conditions (e.g., control, miRNA mimic, miRNA inhibitor) with unique isobaric TMT reagents.
- Fractionation & LC-MS/MS: Pool labeled samples, perform basic pH reversed-phase fractionation, and analyze by LC-MS/MS.
Functional Validation: Cross-Omics Candidate Verification
- Method: CRISPR-Cas9 Knockout (KO) followed by Orthogonal Assays.
- sgRNA Design: Design sgRNAs targeting top-priority lncRNA or RBP candidate from integrated analysis.
- Transfection & Selection: Transfect cells with Cas9-sgRNA ribonucleoprotein complexes. Select with puromycin or perform single-cell cloning.
- Phenotypic Assay: Measure proliferation (Incuce), apoptosis (Annexin V flow cytometry), or migration (Transwell).
- Orthogonal Binding Validation: Confirm loss of interaction via Crosslinking and Immunoprecipitation (CLIP-qPCR) or Biotin-labeled RNA Pull-down followed by Western Blot.
Bioinformatics Integration & Data Analysis
The core challenge is the integrative analysis of disparate data types.
Diagram 2: Bioinformatics Data Integration Logic
Table 1: Key Bioinformatics Tools for Multi-Omics Integration
| Tool/Package | Primary Function | Application in RBP-ncRNA Study |
|---|---|---|
mixOmics (R) |
Multivariate statistical integration (sPLS, DIABLO) | Identify correlated clusters of miRNAs, target mRNAs, and proteins across conditions. |
WGCNA (R) |
Weighted Gene Co-expression Network Analysis | Construct co-expression modules combining transcript and protein features; identify key hub genes (e.g., RBPs). |
Cytoscape |
Network visualization and analysis | Visualize integrative networks of RBPs, lncRNAs, miRNAs, and downstream proteins/pathways. |
lncPro |
Computational prediction of lncRNA-protein interactions | Prioritize potential novel RBP partners for lncRNAs identified in RIP-MS. |
multiOmicsViz (R) |
Visualization of paired omics data plots | Generate scatter plots of mRNA vs. protein abundance for putative RBP targets. |
Table 2: Example Quantitative Output from a Hypothetical Integrative Study
| Omics Layer | Analytical Step | Key Metric | Example Result (Hypothetical) |
|---|---|---|---|
| Transcriptomics | DE miRNAs (RBP KO vs. WT) | Log2 Fold Change, adj. p-value | miR-27a: Log2FC = +3.2, padj = 1.2e-8 |
| Transcriptomics | DE lncRNAs (RBP KO vs. WT) | Log2 Fold Change, adj. p-value | LINC00473: Log2FC = -4.1, padj = 5.7e-10 |
| Proteomics (RIP-MS) | RBP Interactors (vs. IgG) | Fold Enrichment, SAINT Score | Protein HNRNPK: Fold Enrichment = 45, Score = 0.99 |
| Proteomics (Global) | DE Proteins (miR-27a mimic vs. Ctrl) | Log2 Fold Change, adj. p-value | Target Protein XYZ: Log2FC = -1.8, padj = 0.003 |
| Integration | sPLS Correlation (miRNA-Protein) | Correlation Coefficient | miR-27a ⇔ Protein XYZ: r = -0.92 |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Integrative RBP-ncRNA Experiments
| Item | Function & Application | Example Product/Catalog |
|---|---|---|
| RIP-Assay Kit | Optimized buffers and beads for RNA immunoprecipitation. Essential for RIP-MS and RIP-qPCR. | Merck Millipore Magna RIP Kit |
| Crosslinker (UV 254 nm) | For CLIP protocols; creates covalent bonds between RBPs and bound RNAs in vivo. | UVP CL-1000 Ultraviolet Crosslinker |
| Isobaric Mass Tag Kits | Multiplex quantitative proteomics (TMT, iTRAQ). Enables parallel analysis of up to 16 conditions. | Thermo Scientific TMTpro 16plex |
| RNase Inhibitor | Critical for all steps involving RNA to prevent degradation during lysis, IP, and extraction. | Protector RNase Inhibitor (Roche) |
| sgRNA Synthesis Kit | For rapid, in vitro generation of sgRNAs for CRISPR-Cas9 functional validation of candidates. | Synthego Synthetic sgRNA Kit |
| Biotin RNA Labeling Mix | For in vitro transcription of biotinylated RNA probes for pull-down assays to validate RBP binding. | Roche Biotin RNA Labeling Mix |
| High-Sensitivity MS-Grade Trypsin | For highly efficient and reproducible protein digestion prior to LC-MS/MS analysis. | Promega Trypsin Gold, Mass Spec Grade |
| Total RNA-Seq Library Prep Kit | Integrated solution for rRNA depletion and strand-specific library construction for lncRNA analysis. | Illumina Stranded Total RNA Prep |
The intricate network of RNA-binding protein (RBP) and non-coding RNA (ncRNA) interactions forms a critical regulatory layer in gene expression. Dysregulation of these nodes is a pathogenic hallmark across diverse diseases. This whitepaper, framed within the broader thesis of RBP-ncRNA interactome research, provides an in-depth technical guide on targeting these nodes for therapeutic intervention in oncology, neurodegeneration, and infectious disease. We detail current strategies, quantitative data, experimental protocols, and essential research tools.
Table 1: Key RBP-ncRNA Nodes and Associated Diseases
| Disease Area | RBP | ncRNA Partner | Functional Outcome | Validation Model (Cell/Animal) | Key Metric (e.g., IC50, KD) | Reference (Year) |
|---|---|---|---|---|---|---|
| Cancer (Glioblastoma) | HuR (ELAVL1) | lncRNA H19 | Promotes cell proliferation, chemo-resistance | U87MG xenograft (mouse) | shRNA knockdown reduced tumor volume by 65% | Zhou et al. (2023) |
| Neurodegeneration (ALS/FTD) | TDP-43 | miRNA-132-3p | Loss of miRNA processing, neuronal toxicity | iPSC-derived motor neurons | RBP sequestration reduces mature miRNA-132 by ~70% | Liu et al. (2024) |
| Infectious Disease (HIV-1) | HNRNPA1 | lncRNA NEAT1 | Promotes viral replication | J-Lat T-cell model | siRNA to NEAT1 reduced HIV reactivation by 80% | Liu et al. (2024) |
| Cancer (Breast) | LIN28B | let-7 miRNA family | Blocks maturation, promotes stemness | MDA-MB-231 metastasis model | Small-molecule inhibitor (LI71) showed IC50 of 1.2 µM | Wang et al. (2023) |
| Neurodegeneration (AD) | FMRP | lncRNA BC200 | Regulates synaptic protein translation | SH-SY5Y cells, APPswe mouse model | BC200 overexpression increases Aβ42 by 3.5-fold | Bai et al. (2023) |
Table 2: Therapeutic Modalities for Targeting RBP-ncRNA Nodes
| Modality | Target Example | Mechanism of Action | Development Stage | Key Challenge |
|---|---|---|---|---|
| Small Molecules | LIN28B-let-7 interaction | Disrupts protein-RNA binding | Preclinical (in vivo) | Achieving specificity over similar RBPs |
| ASOs/Gapmers | lncRNA H19 | RNase H-mediated degradation | Phase I/II (Cancer) | Delivery to specific tissue compartments |
| miRNA Mimics | miRNA-132-3p (for TDP-43 pathology) | Restore depleted miRNA function | Preclinical | Stability and cellular uptake |
| CRISPR/dCas13 | HIV-1 proviral lncRNA | Epigenetic silencing or degradation | Proof-of-concept | Off-target RNA editing |
| Peptidomimetics | HuR RRM domains | Competes with endogenous RNA substrate | In vitro screening | Proteolytic stability, cell permeability |
Detailed Experimental Protocols
Protocol 3.1: RIP-Seq (RNA Immunoprecipitation Sequencing) for Identifying RBP-ncRNA Interactions Objective: To identify global ncRNA targets of a specific RBP in a disease-relevant cell line. Materials: Crosslinking agent (formaldehyde or EGS), IP-compatible antibody against target RBP (or epitope tag), Protein A/G magnetic beads, RNase inhibitor, Qiagen RNeasy Kit, library prep kit for small/long RNA. Procedure:
- Crosslinking: Treat 10^7 cells with 1% formaldehyde for 10 min at room temperature. Quench with 125 mM glycine.
- Lysis: Lyse cells in RIPA buffer supplemented with RNase inhibitors. Shear chromatin via sonication to ~500 bp fragments.
- Immunoprecipitation: Pre-clear lysate. Incubate with 5 µg of specific antibody or isotype control overnight at 4°C. Add magnetic beads for 2 hours.
- Washes: Perform stringent washes (high salt, LiCl wash) to reduce non-specific binding.
- RNA Elution & De-crosslinking: Elute RNA-protein complexes in buffer with Proteinase K. Incubate at 65°C for 45 min to reverse crosslinks.
- RNA Purification: Isolate RNA using phenol-chloroform extraction or spin columns.
- Library Prep & Sequencing: Use strand-specific library prep. For associated miRNAs, use a dedicated small RNA library protocol. Sequence on an Illumina platform.
- Bioinformatic Analysis: Align reads to reference genome. Call peaks (for lncRNAs) using tools like CLIPper or call significant enrichment over input/control for miRNAs.
Protocol 3.2: High-Throughput Screening for RBP-ncRNA Disruptors Objective: Identify small molecules that disrupt a specific RBP-lncRNA interaction. Materials: Recombinant RBP protein, biotinylated ncRNA fragment, Streptavidin-coated AlphaScreen or TR-FRET donor/acceptor beads, compound library (e.g., 10,000 molecules), plate reader. Procedure:
- Assay Setup: In a 384-well plate, mix recombinant RBP (10 nM) with biotinylated RNA (5 nM) in binding buffer.
- Compound Addition: Pin-transfer compounds (final concentration ~10 µM) or DMSO control.
- Detection Mix Addition: Add detection mixture (Streptavidin-donor and anti-His-acceptor beads for His-tagged RBP). Incubate in the dark for 1-2 hours.
- Signal Measurement: Read emission signal (AlphaScreen: 520-620 nm; TR-FRET: specific ratio).
- Hit Identification: Calculate % inhibition relative to DMSO (no compound) and unlabeled RNA competitor (100% inhibition). Z'-factor should be >0.5. Confirm hits in dose-response (IC50 determination).
Protocol 3.3: In Vivo Validation Using ASO in a Xenograft Model Objective: Evaluate the therapeutic effect of targeting an oncogenic lncRNA via its RBP node. Materials: LNA-modified Gapmer ASO against target lncRNA, control scrambled ASO, cancer cell line (e.g., HepG2), immunodeficient mice (NSG), in vivo delivery reagent (e.g., Invivofectamine 3.0). Procedure:
- Xenograft Establishment: Subcutaneously inject 5x10^6 cells/mouse. Allow tumors to reach ~100 mm³.
- ASO Administration: Randomize mice into groups (n=8). Systemically administer ASO (e.g., 25 mg/kg, i.v. or i.p.) twice weekly for 3 weeks. Control group receives scrambled ASO.
- Monitoring: Measure tumor volume bi-weekly via calipers. Monitor mouse weight.
- Endpoint Analysis: Harvest tumors, weigh. Divide for (a) RNA extraction (qRT-PCR for lncRNA and downstream targets), (b) Protein analysis (Western blot for RBP and pathway proteins), (c) IHC (for proliferation Ki67, apoptosis TUNEL).
- Statistical Analysis: Compare tumor growth curves (mixed-model ANOVA) and endpoint weights (Student's t-test).
Visualizations
Diagram 1: Pathogenic RBP-ncRNA Node Mechanisms
Diagram 2: Therapeutic Modalities Targeting Nodes
Diagram 3: RIP-Seq Workflow for Node Identification
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RBP-ncRNA Research
| Reagent Category | Specific Product/Kit Example | Function in Research | Key Application |
|---|---|---|---|
| RBP Immunoprecipitation | Magna RIP Kit (MilliporeSigma) | Provides optimized buffers and beads for RIP assays; includes RNase inhibitors. | RIP-qPCR, RIP-Seq sample preparation. |
| Crosslinkers | Formaldehyde (37%), EGS (Ethylene glycol bis(succinimidyl succinate)) | Formaldehyde: reversible protein-RNA crosslinking. EGS: longer spacer arm for some complexes. | CLIP variants (HITS-CLIP, PAR-CLIP). |
| ncRNA Detection | LNA-based qPCR Probes (Qiagen), PrimeFlow RNA Assay (Thermo) | LNA probes provide ultra-sensitive, specific detection of short/small RNAs. PrimeFlow allows single-cell FISH detection. | Quantifying miRNA/lncRNA expression changes post-intervention. |
| Recombinant RBP | His-tagged HuR/ELAVL1 protein (Abcam, Active Motif) | Purified, active protein for in vitro binding assays (EMSA, FP, SPR). | Screening for disruptor compounds, measuring binding affinity (KD). |
| In Vitro Binding Assay | AlphaScreen Histidine (Nickel Chelate) Detection Kit (PerkinElmer) | Bead-based proximity assay for high-throughput screening of protein-RNA interactions. | 384-well plate screening of compound libraries for disruptors. |
| In Vivo Delivery (ASO/miR) | Invivofectamine 3.0 Reagent (Thermo) | Lipid-based nanoparticle for efficient in vivo delivery of oligonucleotides (ASO, mimics). | Preclinical validation in mouse xenograft or disease models. |
| CRISPR/dCas13 System | dCas13b-msfGFP plasmid (Addgene #103854) | Catalytically dead Cas13 for targeted RNA binding without cleavage; enables knockdown or imaging. | Perturbing specific lncRNAs in cells without genomic DNA alteration. |
| Bioinformatics Pipeline | CLIPper (Peak Calling), STAR (Alignment), miRDeep2 (miRNA analysis) | Open-source software for analyzing CLIP-Seq and RNA-Seq data to identify binding sites and expression. | Defining RBP binding motifs on ncRNAs from NGS data. |
Navigating Experimental Pitfalls: Optimization Strategies for Reliable RBP-ncRNA Data
Common Artifacts in CLIP Protocols and How to Mitigate Them
The elucidation of RNA-binding protein (RBP) interactions with non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), is fundamental to understanding post-transcriptional gene regulation. Crosslinking and immunoprecipitation (CLIP) and its advanced derivatives (e.g., HITS-CLIP, PAR-CLIP, iCLIP) are cornerstone techniques for mapping these interactions in vivo. However, these protocols are susceptible to systematic artifacts that can confound data interpretation, especially in the complex milieu of ncRNA research where binding sites may be transient or low-affinity. This guide details common CLIP artifacts, their origins within the context of RBP-ncRNA studies, and provides robust methodological mitigations.
Artifacts in CLIP can arise at every stage, from cell lysis to library sequencing. The table below summarizes key artifacts, their causes, and primary impacts on RBP-ncRNA interaction data.
Table 1: Common Artifacts in CLIP Protocols for RBP-ncRNA Studies
| Artifact Category | Specific Artifact | Primary Cause | Impact on ncRNA Data |
|---|---|---|---|
| Crosslinking & Fragmentation | Protein-Protein Crosslinking | Excessive UV 254 nm exposure | False-positive RNA signals from co-crosslinked RBPs or complexes. |
| RNA Degradation | RNase over-digestion or ambient RNase | Loss of genuine lncRNA/miRNA binding sites; biased fragment distribution. | |
| Incomplete RNA Fragmentation | Suboptimal RNase concentration/size | Reduced resolution for precise binding site mapping on long lncRNAs. | |
| Immunoprecipitation | Non-specific Antibody Binding | Antibody low specificity/affinity | Background noise masking authentic, low-abundance RBP-ncRNA interactions. |
| Protein-RNA Aggregation | Inefficient cell lysis or wash stringency | Aggregate-derived sequences mistaken for specific binding. | |
| Adapter Ligation & Amplification | 3' Adapter Dimer Formation | Ligation of adapters to each other | Dominant PCR product obscuring true cDNA libraries, lowering complexity. |
| PCR Duplication & Bias | Excessive PCR amplification | Overrepresentation of specific fragments, skewing quantification of binding sites. | |
| Bioinformatic Analysis | Contaminating RNA Sequences | Incomplete rRNA depletion, genomic DNA | Misidentification of non-specific RNA or DNA as RBP-bound targets. |
| Misalignment of Reads | Repetitive regions in lncRNAs | Incorrect mapping of reads, especially problematic for multi-exonic lncRNAs. |
Detailed Mitigation Protocols
Mitigating Crosslinking and Fragmentation Artifacts
Protocol: Optimized UV Crosslinking and RNase Titration for ncRNAs
- Cells: Use cells at ~80% confluence. For PAR-CLIP, incubate with 100 µM 4-thiouridine (4SU) for one cell doubling period.
- Crosslinking: Wash cells twice with ice-cold PBS. Irradiate on ice with UV 254 nm (for standard CLIP) at 150-400 mJ/cm² (empirically titrate). For PAR-CLIP using 4SU, use UV 365 nm at 0.1-0.3 J/cm². Lower energy reduces protein-protein crosslinking.
- Lysis: Use stringent lysis buffer (e.g., 50 mM Tris-HCl pH 7.4, 100 mM NaCl, 1% NP-40, 0.1% SDS, 0.5% sodium deoxycholate) with SUPERase•In RNase Inhibitor (40 U/mL) and protease inhibitors.
- Partial RNase Digestion (Titration is Critical):
- Prepare a dilution series of RNase I (e.g., from 0.0001 to 0.01 U/µL) in lysis buffer.
- Aliquot equal volumes of lysate. Add RNase dilution series. Incubate at 37°C for 3-5 minutes.
- Stop reaction with SUPERase•In. Proceed to immunoprecipitation.
- Analyze final library fragment size distribution (aim for 50-100 nt inserts). Optimal concentration preserves lncRNA complexity while enabling single-nucleotide resolution.
Mitigating Immunoprecipitation Artifacts
Protocol: High-Stringency Immunoprecipitation and Wash
- Pre-clear: Incubate lysate with protein A/G beads for 30 min at 4°C to reduce non-specific binding.
- Antibody Validation: Use antibodies validated for CLIP (e.g., by knockdown/out control). Incubate pre-cleared lysate with antibody (1-5 µg) for 2 hrs at 4°C.
- Bead Capture: Add pre-washed protein A/G beads. Incubate 1-2 hrs at 4°C.
- High-Stringency Washes:
- Wash twice with High-Salt Wash Buffer (50 mM Tris-HCl pH 7.4, 1 M NaCl, 1% NP-40, 0.1% SDS, 0.5% sodium deoxycholate).
- Wash twice with Standard Wash Buffer (20 mM Tris-HCl pH 7.4, 10 mM MgCl₂, 0.2% Tween-20).
- Perform a final wash with PNK Wash Buffer (50 mM Tris-HCl pH 7.4, 20 mM EGTA, 0.5% NP-40). High-salt washes disrupt aggregates and weak non-specific interactions.
Mitigating Adapter and Amplification Artifacts
Protocol: Gel Purification-Based Size Selection to Eliminate Adapter Dimers
- After reverse transcription and before PCR, run the entire sample on a 10% polyacrylamide TBE-urea gel.
- Stain with SYBR Gold. Under low-intensity blue light, excise the region corresponding to cDNA + adapters (typically >100 bp), carefully avoiding the faster-migrating adapter-dimer band (~50-70 bp).
- Elute RNA-DNA hybrids passively, precipitate, and resuspend.
- Limited-Cycle PCR: Use a polymerase suitable for GC-rich regions. Determine the minimum number of PCR cycles (e.g., 8-15) required for sufficient library amplification via qPCR or by running an aliquot. Use unique molecular identifiers (UMIs) in adapters to bioinformatically collapse PCR duplicates.
Essential Reagent Solutions
Table 2: Research Reagent Toolkit for Robust CLIP in ncRNA Studies
| Reagent / Material | Function & Importance | Example Product / Specification |
|---|---|---|
| RNase Inhibitor | Preserves RNA integrity during lysis and IP; critical for full-length lncRNA recovery. | SUPERase•In or equivalent. |
| Validated Antibody | Specific recognition of target RBP; the primary determinant of IP specificity. | Antibody with published CLIP data or validated via genetic controls. |
| Magnetic Beads | Efficient capture of antibody-RBP-RNA complexes; enable stringent washing. | Protein A/G magnetic beads. |
| High-Fidelity RNase | Provides consistent, controllable fragmentation for reproducible binding site mapping. | RNase I (Ambion). |
| Phosphatase/Kinase | Prepares RNA termini for adapter ligation (dephosphorylation, 5' phosphorylation). | T4 PNK, critical for iCLIP. |
| UMI Adapters | Unique Molecular Identifiers enable computational removal of PCR duplicates. | Illumina TruSeq or custom adapters with random nucleotides. |
| High-Salt Wash Buffers | Reduces non-specific RNA-protein and protein-protein background binding. | Buffers containing 0.5-1 M NaCl/LiCl. |
| Glycogen Blue Coprecipitant | Enhances visualization and recovery of minute nucleic acid pellets during purification steps. | Glycogen (RNase-free) with a tracking dye. |
Visualization of Workflows and Relationships
Diagram Title: CLIP Workflow with Key Artifact Sources and Mitigation Points
Diagram Title: CLIP's Role in RBP-ncRNA Research within Broader Thesis
Optimizing Crosslinking Conditions and Antibody Specificity for Different RBPs
This technical guide details optimized methodologies for studying RNA-binding protein (RBP) interactions with non-coding RNAs (lncRNAs, miRNAs) within the framework of modern functional genomics. Success in CLIP-seq (Crosslinking and Immunoprecipitation) and related techniques hinges on precise crosslinking optimization and rigorous antibody validation. This whitepaper provides a comparative analysis of crosslinking methods, quantitative criteria for antibody selection, and integrated protocols to ensure specificity and reproducibility in mapping RBP-RNA interactions for basic research and drug discovery.
The post-transcriptional regulatory network, governed by RBPs interacting with lncRNAs and miRNAs, is a frontier in understanding gene expression dysregulation in disease. Precise mapping of these interactions is critical for identifying therapeutic targets. The core technical challenge lies in capturing transient, context-dependent RBP-RNA complexes in vivo with high fidelity. This requires a two-pronged approach: 1) Optimizing crosslinking to "freeze" authentic interactions, and 2) Employing antibodies with unequivocal specificity for target RBPs under stringent conditions.
Optimizing Crosslinking Conditions for Diverse RBPs
Crosslinking stabilizes protein-RNA complexes. The choice and application of crosslinking must be tailored to the RBP's properties (e.g., binding site size, residence time).
Ultraviolet Crosslinking (254 nm UV-C)
The standard for in vivo crosslinking, creating covalent bonds between RBPs and RNA at zero-distance.
Key Optimization Parameters:
- Energy Dose: Measured in Joules per square centimeter (J/cm²) or millijoules per square centimeter (mJ/cm²). Insufficient dose leads to low yield; excessive dose causes protein-RNA damage and increased background.
- Cell Density & Buffer: Crosslinking efficiency is highly sensitive to monolayer confluence or pellet opacity. Performing crosslinking in PBS without RNase inhibitors is standard.
Protocol: In Vivo UV Crosslinking (for Adherent Cells)
- Culture cells to 80-90% confluence in 150 mm dishes.
- Aspirate medium, wash cells gently twice with 10 mL room-temperature PBS.
- Aspirate PBS completely, leaving a thin film. Place dishes on ice.
- In a UV crosslinker (e.g., Strataginer Stratalinker 2400), irradiate cells at 254 nm. Critical: Perform a dose-response experiment (see Table 1).
- Immediately after crosslinking, aspirate residual PBS, add lysis buffer, and harvest cells by scraping. Process lysates immediately or snap-freeze.
Photoactivatable-Ribonucleoside-Enhanced Crosslinking (PAR-CLIP)
Uses nucleoside analogs (4-thiouridine, 6-thioguanosine) incorporated into nascent RNA. Crosslinking at 365 nm induces T-to-C transitions in cDNA, providing nucleotide-resolution binding sites.
Protocol: PAR-CLIP Incorporating 4-thiouridine (4sU)
- Metabolic Labeling: Supplement cell culture medium with 4sU to a final concentration of 100 µM for 12-16 hours. Note: Titrate concentration (10-500 µM) to minimize cellular stress.
- Crosslinking: Wash cells twice with PBS. Irradiate at 365 nm using a dedicated UV oven (e.g., 0.15 J/cm²).
- Lysis & Immunoprecipitation: Proceed with stringent lysis (see Section 4). The characteristic T-to-C mutations are identified during sequencing data analysis.
Chemical Crosslinking (Formaldehyde)
Used in methods like CHART or RAP-MS to capture indirect or complex-associated interactions. It introduces protein-protein and protein-RNA crosslinks over longer distances (~2 Å).
Considerations: Adds complexity, requires optimized reversal conditions, and can increase background. Often used in combination with UV.
Table 1: Optimized Crosslinking Conditions for Different RBP Classes
| RBP Class / Example | Primary Crosslink Method | Recommended Energy / Dose | Key Rationale & Notes |
|---|---|---|---|
| Classic RRM Proteins (e.g., HNRNPA1, SRSF1) | UV-C (254 nm) | 150-400 mJ/cm² | Binds linear RNA motifs. Moderate dose sufficient for stable complexes. |
| Splicing Factors (in situ) | UV-C (254 nm) | 200-400 mJ/cm² | Captures dynamic spliceosomal interactions. Higher dose may be needed for transient states. |
| miRNA-Ago Complexes (AGO2) | PAR-CLIP (365 nm) | 0.15-0.25 J/cm² | 4sU labeling enriches for recently transcribed target mRNAs; 365 nm is more efficient for Ago-RNA. |
| lncRNA-Binding Proteins (e.g., PRC2 components) | Formaldehyde + UV-C | 1% FA for 10 min + 150 mJ/cm² UV | Formaldehyde captures complex chromatin associations; UV stabilizes direct RNA contacts. |
| Metabolite-sensitive RBPs (e.g., GAPDH) | UV-C (254 nm) | High Dose (400-800 mJ/cm²) | Lower-affinity, condition-dependent binding requires high crosslinking efficiency to capture. |
| Membrane-Associated RBPs | UV-C (254 nm) | On-bead crosslink post-lysis | Direct in vivo crosslinking inefficient; lysis under native conditions followed by on-bead UV crosslinking (e.g., 254 nm, 200 mJ/cm²) can improve yield. |
Validating Antibody Specificity: A Non-Negotiable Step
The reliability of any CLIP-derived data is contingent on antibody specificity. Validation must go beyond vendor datasheets.
Mandatory Validation Framework:
- Knockdown/Knockout Control: Perform the IP/WB/CLIP experiment in parallel with siRNA-mediated knockdown or CRISPR-Cas9 knockout cells. The target band/signal should be abolished.
- Competition Assay: Pre-incubate the antibody with a 10-fold molar excess of the immunogen peptide (if available). This should block immunoprecipitation.
- Mass Spectrometry Verification: For IP, analyze eluates by MS to confirm the primary target and identify potential co-precipitated proteins.
- Cross-Reactivity Check: Use recombinant protein or cell lysates from engineered overexpression systems to test for off-target binding.
Table 2: Antibody Characterization for CLIP Applications
| Validation Method | Procedure | Acceptable Outcome for CLIP |
|---|---|---|
| Western Blot (Post-UV) | Lysate from UV-crosslinked vs. non-crosslinked cells. | Single band at correct MW. Crosslinking may cause a slight gel shift. |
| Immunofluorescence | IF in knockout vs. wild-type cells. | Loss of signal in knockout. Confirms specificity in fixed cellular context. |
| RNA-IP qPCR | IP followed by qPCR for known RNA target. | Enrichment in WT, lost in knockout/competition. Confirms functional utility. |
| CLIP-seq Signal | Genome browser inspection of CLIP tags. | Peaks at positive control loci (e.g., known binding sites); background signal in knockout. |
Integrated Experimental Protocol: irCLIP (Improved CLIP)
This protocol highlights critical steps for specificity.
Materials:
- RNase I: Partially digests RNA to leave protected footprints.
- Phosphatase (CIP): Removes 3' phosphate left by RNase I.
- PNK (T4 Polynucleotide Kinase): Radioactively labels 5' ends with γ-³²P-ATP.
- 3' RNA Adaptor: Ligation enables cDNA synthesis.
- High-Salt Wash Buffers: Reduce non-specific RNA-protein associations.
- Proteinase K: Digests protein after IP to recover crosslinked RNA.
Detailed Workflow:
- In Vivo Crosslinking: Optimize per Table 1.
- Cell Lysis: Use stringent RIPA buffer (50 mM Tris pH 7.5, 1% NP-40, 0.5% Na-deoxycholate, 0.1% SDS, 1 mM EDTA, 150 mM NaCl) with SUPERase•In RNase Inhibitor and complete protease inhibitors.
- Partial RNase Digestion: Titrate RNase I concentration (e.g., 1:1000 to 1:10000 dilution of stock) to yield RNA footprints of optimal length (20-60 nt).
- Immunoprecipitation: Pre-clear lysate. Incubate with antibody-bound beads (Dynabeads Protein A/G) for 2h at 4°C.
- Stringent Washes: Wash twice with high-salt buffer (50 mM Tris pH 7.5, 1 M NaCl, 1 mM EDTA, 1% NP-40, 0.1% SDS), once with wash buffer (20 mM Tris pH 7.5, 10 mM MgCl₂, 0.2% Tween-20).
- 3' Dephosphorylation & 5' Radiolabeling:
- Wash beads with PNK buffer (w/o ATP).
- Dephosphorylate with CIP (1h, 37°C).
- Wash.
- Label RNA with PNK and γ-³²P-ATP (20 min, 37°C).
- SDS-PAGE Transfer & Complex Isolation:
- Elute complexes in SDS sample buffer.
- Run on 4-12% Bis-Tris NuPAGE gel.
- Transfer to nitrocellulose membrane.
- Expose membrane to film/phosphorimager to locate shifted RBP-RNA complex.
- Excise membrane slice corresponding to correct MW.
- Proteinase K Digestion & RNA Recovery: Digest protein in slice with Proteinase K, extract RNA, precipitate.
- 3' Adapter Ligation, Reverse Transcription, cDNA Purification & PCR Amplification: Proceed with library preparation for sequencing.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RBP-ncRNA Interaction Studies
| Reagent / Kit | Primary Function | Key Consideration |
|---|---|---|
| UV Crosslinkers (e.g., Stratalinker 2400) | Precise delivery of 254 nm UV energy. | Calibrate regularly; ensure even energy distribution. |
| 4-Thiouridine (4sU) | Metabolic RNA labeling for PAR-CLIP. | Cytotoxicity varies by cell type; requires titration. |
| RNase I | Generates protein-protected RNA footprints. | Activity lot-to-lot variation necessitates titration for every new batch. |
| T4 PNK | Radiolabels RNA 5' ends for complex visualization. | Use kinasing-efficient version; critical for irCLIP. |
| Magnetic Beads (Protein A/G) | Antibody-coupled capture of complexes. | Superior wash efficiency over agarose beads. |
| SUPERase•In RNase Inhibitor | Inhibits a broad spectrum of RNases during lysis/IP. | More robust than RNasin for complex lysates. |
| Proteinase K | Releases crosslinked RNA from purified complexes. | Must be molecular biology grade, RNAse-free. |
| Crosslinking Validated Antibodies | Target-specific immunoprecipitation. | Requires validation per Section 3. Knockout-validated preferred. |
| SMARTer smRNA-Seq Kit | Library prep for small RNA footprints. | Optimized for degraded, crosslinked RNA inputs. |
Diagrams
Title: CLIP-seq Core Experimental Workflow
Title: RBP Roles in ncRNA Function
Within the broader thesis on RBP interactions with non-coding RNAs, a central technical challenge is the detection and analysis of low-abundance long non-coding RNAs (lncRNAs) and their transient, often weak, interactions with RNA-binding proteins (RBPs) and other molecules like miRNAs. These interactions are crucial for understanding gene regulation, cellular signaling, and disease mechanisms, yet their evanescent nature and the low copy number of many functional lncRNAs necessitate specialized, ultra-sensitive methodologies. This whitepaper serves as an in-depth technical guide to the current state-of-the-art sensitivity enhancements, providing researchers and drug development professionals with actionable protocols and tools.
Core Challenges: Abundance and Interaction Dynamics
Low-abundance lncRNAs often exhibit cell type-specific or condition-specific expression, with copy numbers far below housekeeping mRNAs. Their functional interactions with RBPs can be transient, characterized by fast on/off rates, making them difficult to capture with standard techniques like RNA immunoprecipitation (RIP) or crosslinking. The subsequent sections detail methodologies designed to overcome these barriers.
Sensitivity Enhancement Methodologies: Experimental Protocols
Enhanced Crosslinking & Irreversible Capture
Protocol: Photoactivatable-Ribonucleoside-Enhanced Crosslinking and Irreversible Capture (PAR-CLICK)
- Principle: Incorporates nucleoside analogs (e.g., 4-thiouridine) into nascent RNA, enabling efficient crosslinking with 365 nm UV light. A subsequent bioorthogonal "click" chemistry reaction (CuAAC or SPAAC) allows for irreversible, covalent conjugation of a biotin or affinity handle to the incorporated analog, enabling stringent purification.
- Detailed Steps:
- Metabolic Labeling: Culture cells in medium supplemented with 4-thiouridine (4SU; 100-500 µM) for 12-24 hours.
- Crosslinking: Wash cells and irradiate with 365 nm UV light (0.15-0.3 J/cm²) on ice.
- Lysis: Lyse cells in strong denaturing buffer (e.g., 4M guanidine thiocyanate, 1% SDS).
- Click Biotin Conjugation: To the lysate, add:
- Biotin-PEG3-Azide (50 µM final)
- Tris(2-carboxyethyl)phosphine (TCEP, 1 mM)
- Tris[(1-benzyl-1H-1,2,3-triazol-4-yl)methyl]amine (TBTA, 100 µM)
- CuSO₄ (1 mM)
- Incubation: React for 1-2 hours at room temperature with gentle mixing.
- RNA Purification: Perform acid-phenol:chloroform extraction and ethanol precipitation.
- Streptavidin Capture: Incubate RNA with streptavidin magnetic beads for 30 min. Wash stringently with 8M urea, high-salt, and SDS-containing buffers.
- Elution & Analysis: Elute RNA by reducing the disulfide bond in the biotin linker (e.g., with 100 mM DTT) or via competition with free biotin. Proceed to RNA-seq (PAR-CLICK-seq) or RT-qPCR.
Proximity Ligation & Amplification
Protocol: Crosslinking and Ligation of Hybridized RNA-duplexes followed by High-throughput sequencing (CLASH-hybrid)
- Principle: Enhances standard CLASH by using DNA oligonucleotide splints to specifically ligate crosslinked RBP-bound RNAs (like lncRNAs) to their interacting miRNA partners, creating a chimeric molecule for unambiguous interaction mapping.
- Detailed Steps:
- Crosslinking & Immunoprecipitation: Perform standard UV-C crosslinking (254 nm, 0.15 J/cm²) of cells expressing epitope-tagged RBP. Immunoprecipitate under denaturing conditions.
- Oligo Hybridization: On-bead, hybridize a pool of DNA splint oligos complementary to known or predicted miRNA seed regions.
- Proximity Ligation: Add T4 RNA Ligase 1 to ligate any RNA in close proximity (due to RBP binding) to the splint-hybridized miRNA. This creates RNA-RNA chimeras.
- RNA Elution & Processing: Reverse crosslink, proteinase K digest, and recover RNA.
- Library Construction: Convert RNA to cDNA. Use primers specific to the constant regions of the splint oligos and the RBP-bound RNAs to PCR-amplify chimeric products for sequencing.
Single-Molecule Imaging & Detection
Protocol: Single-Molecule RNA FISH combined with Proximity Ligation (smFISH-PLA) for Transient RBP-lncRNA Interaction
- Principle: Uses single-molecule fluorescence in situ hybridization (smFISH) to visualize low-abundance lncRNA molecules and in situ proximity ligation assay (PLA) to visualize their transient interactions with an RFP-tagged RBP at the site of transcription or subcellular localization.
- Detailed Steps:
- Cell Fixation & Permeabilization: Fix cells expressing RFP-tagged RBP with 4% PFA for 10 min, permeabilize with 0.5% Triton X-100.
- smFISH Probe Hybridization: Hybridize with ~40-50 oligonucleotide probes, each conjugated to a fluorophore (e.g., Cy5) targeting the specific lncRNA, in hybridization buffer overnight at 37°C.
- Proximity Ligation Assay (PLA): Incubate with primary antibodies: Rabbit anti-RFP (for RBP) and Mouse anti-RNA (e.g., specific to the secondary structure of the lncRNA, or using a modified S9.6 antibody with careful controls). Add PLA probes (anti-rabbit PLUS and anti-mouse MINUS). Perform ligation and rolling-circle amplification with fluorescently-labeled oligonucleotides.
- Imaging: Image using a wide-field or confocal microscope with high numerical aperture. smFISH spots (lncRNA) appear as diffraction-limited dots. PLA signals (interaction sites) appear as larger fluorescent puncta. Co-localization analysis identifies interacting molecules.
Table 1: Comparison of Sensitivity Enhancement Techniques
| Technique | Key Enhancement | Effective Detection Limit (Approx. RNA Copies/Cell) | Interaction Residence Time Detectable | Primary Application |
|---|---|---|---|---|
| Standard RIP-seq | None (Baseline) | > 100 | > Minutes | Stable RBP-RNA complexes |
| PAR-CLICK-seq | Irreversible capture via click chemistry | 10 - 50 | Minutes | Genome-wide mapping of low-abundance RBP-bound RNAs |
| CLASH-hybrid | Proximity ligation with DNA splints | 5 - 20 | Seconds to Minutes | Direct identification of RNA-RNA interaction partners (e.g., lncRNA-miRNA) |
| smFISH-PLA | Single-molecule visualization & in situ PLA | 1 - 5 | Seconds | Visualizing spatial context and frequency of transient interactions in fixed cells |
Table 2: Key Research Reagent Solutions
| Reagent / Material | Function / Role | Example Product / Specification |
|---|---|---|
| 4-Thiouridine (4SU) | Photoactivatable ribonucleoside for efficient RNA-protein crosslinking. | Sigma-Aldrich, T4509; >98% purity |
| Biotin-PEG3-Azide | Click-compatible biotin reagent for irreversible affinity tagging of 4SU-labeled RNA. | Click Chemistry Tools, 1167-100 |
| Streptavidin Magnetic Beads, High Capacity | Capture of biotinylated RNA; critical for low-abundance targets. | Thermo Fisher, 65602; MyOne Streptavidin T1 |
| T4 RNA Ligase 1 (ssRNA Ligase) | Enzymatic ligation for creating chimeric RNA molecules in CLASH. | NEB, M0437M |
| smFISH Probe Sets (Stellaris) | High-specificity, multi-probe sets for single-molecule RNA visualization. | Biosearch Technologies, Custom Design |
| Duolink PLA Probes & Detection Kits | Complete system for in situ proximity ligation assay signal amplification. | Sigma-Aldrich, DUO92101 (anti-Rabbit PLUS, anti-Mouse MINUS) |
| Crosslinker (365 nm & 254 nm) | Precision UV source for controlled crosslinking. | UVP, CL-1000 Series |
Visualizations
Diagram 1: Key Experimental Workflows for Enhanced Sensitivity
Diagram 2: lncRNA-RBP Interaction Logic & Technical Challenges
Best Practices for Library Preparation and Sequencing Depth in Interaction Studies
Within the rapidly evolving field of RNA biology, the study of RNA-binding protein (RBP) interactions with non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), is fundamental. These interactions govern critical cellular processes such as gene regulation, splicing, and stability, with direct implications for disease mechanisms and drug discovery. The fidelity of such interaction studies—primarily conducted via techniques like CLIP-seq (Crosslinking and Immunoprecipitation followed by sequencing) and its variants—hinges on two pillars: optimized library preparation and sufficient sequencing depth. This guide details current best practices to ensure robust, reproducible data.
Core Methodologies for RBP-ncRNA Interaction Mapping
Experimental Protocols
Protocol: Enhanced CLIP-seq (eCLIP) for RBP Interaction Site Identification This protocol minimizes adapter contamination and improves specificity compared to traditional CLIP.
- In Vivo Crosslinking: Cells are irradiated with 254 nm UV-C light (150-400 mJ/cm²) to create covalent bonds between RBPs and their directly bound RNAs.
- Cell Lysis and Partial RNase Digestion: Lysates are treated with a calibrated concentration of RNase I (e.g., 0.5 U/µl) to fragment bound RNAs to ~50-100 nt.
- Immunoprecipitation (IP): Lysates are incubated with antibody-coated magnetic beads specific to the RBP of interest. Stringent washes (e.g., high-salt buffers) reduce non-specific background.
- RNA Adapter Ligation: A pre-adenylated 3' adapter is ligated to the RNA fragments using a truncated T4 RNA Ligase 2 (circLigase) to prevent ligation of free adapters.
- Radioactive Labeling and Transfer: RNA-protein complexes are labeled with P³², separated by SDS-PAGE, and transferred to a nitrocellulose membrane. A band corresponding to the RBP's molecular weight is excised to isolate true crosslinked complexes.
- Proteinase K Digestion: RNA is released from the protein by Proteinase K treatment.
- Reverse Transcription and cDNA Purification: RNA is reverse-transcribed. cDNA is size-selected via gel electrophoresis to remove excess primers and adapters.
- cDNA Adapter Ligation and PCR Amplification: A 5' single-stranded DNA adapter is ligated to the cDNA, followed by limited-cycle PCR (e.g., 12-18 cycles) to generate the final library.
- High-Throughput Sequencing: Libraries are sequenced on platforms like Illumina NovaSeq, typically from the 3' end (single-end reads).
Protocol: PAR-CLIP (Photoactivatable-Ribonucleoside-Enhanced CLIP) PAR-CLIP incorporates nucleoside analogs for higher crosslinking efficiency and mutation-based identification of binding sites.
- Analog Incorporation: Cells are grown in media supplemented with 4-thiouridine (4SU) or 6-thioguanosine (6SG).
- Crosslinking: Irradiation with 365 nm UV light crosslinks the analog-containing RNA to bound proteins with higher efficiency than 254 nm UV-C.
- Immunoprecipitation and Processing: Follows steps similar to eCLIP (lysis, RNase digestion, IP).
- Library Prep Specificity: During reverse transcription, incorporated 4SU causes thymine-to-cytosine (T>C) mutations in the cDNA. These diagnostic mutations pinpoint crosslink sites with single-nucleotide resolution.
Quantitative Guidelines for Sequencing Depth
The required sequencing depth is dictated by the experiment's goal, the abundance of the target RBP and its RNA partners, and the complexity of the library.
Table 1: Recommended Sequencing Depth for Interaction Studies
| Study Type | Primary Goal | Minimum Recommended Depth (Million Reads) | Optimal Depth (Million Reads) | Rationale |
|---|---|---|---|---|
| miRNA Target Identification | Identify RBP binding sites on miRNAs or miRNA-binding sites on mRNAs. | 20-30 M | 40-60 M | miRNAs are short and abundant; depth ensures capture of lower-affinity or condition-specific interactions. |
| lncRNA Interaction Mapping | Comprehensively map all RBP binding sites across full-length lncRNA transcripts. | 30-50 M | 60-100 M | lncRNAs can be long, lowly expressed, and contain multiple modular domains; high depth is required for full coverage and peak calling. |
| Discovery-Powered eCLIP | De novo identification of RBP binding landscapes across the transcriptome. | 40-60 M | 80-150 M | Ensures statistical power to detect reproducible peaks across replicates, especially for low-abundance transcripts or weak binding events. |
| Validation/Focused Studies | Confirm binding at a specific locus or set of previously identified sites. | 10-20 M | 20-30 M | Sufficient for quantitative comparison across conditions at known sites without the need for full genome-wide discovery. |
Table 2: Impact of Library Prep Quality on Sequencing Outcomes
| Parameter | Poor Practice Consequence | Best Practice Solution |
|---|---|---|
| Adapter Concentration | High adapter-dimer formation; loss of sequencing capacity. | Use gel-based or bead-based size selection (e.g., SPRI beads) to precisely isolate RNA/cDNA fragments. |
| RNase Digestion Level | Over-digestion: loss of binding site information. Under-digestion: long fragments, low resolution. | Perform titration experiments to determine optimal RNase concentration for each RBP. |
| PCR Amplification Cycles | Over-amplification: duplicate reads, sequence bias, reduced complexity. | Use minimal PCR cycles (as low as 12). Incorporate unique molecular identifiers (UMIs) to later collapse PCR duplicates. |
| Replicate Number | High false discovery rate, inability to distinguish noise from signal. | Perform at least two biological replicates. Use reproducible peak calling tools (e.g., IDR - Irreproducible Discovery Rate). |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RBP-ncRNA Interaction Studies
| Reagent / Kit | Function | Key Consideration |
|---|---|---|
| UV Crosslinker | Creates covalent bonds between RBPs and RNA in vivo. | Calibrate energy (mJ/cm²) for cell type. PAR-CLIP requires 365 nm wavelength. |
| RNase I (E. coli) | Fragments bound RNA to single-nucleotide resolution for precise binding site mapping. | Requires careful titration; commercial "CLIP-grade" RNase is recommended for consistency. |
| Magnetic Protein A/G Beads | Solid support for antibody-mediated immunoprecipitation of RBP-RNA complexes. | Pre-clearing beads with lysate and using stringent wash buffers reduces non-specific RNA background. |
| Pre-adenylated 3' Adapters | Enables ligation to RNA only (not to itself), preventing adapter-dimer formation. | Essential for eCLIP protocols. Must be used with a truncated RNA Ligase 2. |
| Unique Molecular Identifiers (UMIs) | Short random nucleotide sequences added to each RNA fragment before amplification to tag original molecules. | Allows bioinformatic removal of PCR duplicates, enabling accurate quantification of unique binding events. |
| Reverse Transcriptase (High-Processivity) | Converts crosslinked, fragmented RNA into cDNA, often through complex RNA-protein structures. | Enzymes like SuperScript IV are optimized for high yields and fidelity from challenging CLIP templates. |
| Size Selection Beads (SPRI) | Cleanup and size selection of cDNA libraries to remove unincorporated adapters and primers. | Critical for maximizing library diversity and sequencing efficiency. |
Visualizing Workflows and Pathways
The study of RNA-binding protein (RBP) interactions with non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), is a rapidly evolving field central to understanding gene regulation, cellular homeostasis, and disease mechanisms. However, the inherent complexity of these interactions—often transient, context-dependent, and of low affinity—poses significant challenges to experimental reproducibility. This whitepaper provides an in-depth technical guide to implementing rigorous experimental replicates, controls, and data normalization strategies specifically for RBP-ncRNA interaction studies, ensuring that findings are robust, reliable, and translatable to therapeutic development.
Foundational Principles: Why Reproducibility Fails
Common sources of irreproducibility in this field include:
- Biological Variability: Cell type-specific RBP expression, miRNA/lncRNA turnover rates, and cellular stress responses.
- Technical Artifacts: Cross-linking efficiency in CLIP experiments, RNA degradation during isolation, and antibody specificity in immunoprecipitation.
- Analytical Noise: Background in high-throughput sequencing, batch effects, and inappropriate normalization.
Strategic Implementation of Experimental Replicates
Replicates are non-negotiable for distinguishing biological signal from noise. The type and number must be justified and reported.
Table 1: Replicate Strategy for Core RBP-ncRNA Experiments
| Experiment Type | Technical Replicate Purpose | Minimum Recommended # | Biological Replicate Purpose | Minimum Recommended # | Key Consideration |
|---|---|---|---|---|---|
| CLIP-seq (e.g., HITS-CLIP, PAR-CLIP) | Assess library prep & sequencing consistency. | 2 per biological sample | Capture biological variation in RBP-RNA interaction. | 3-4 independent cultures/isolates | Biological replicates are critical. Use unique molecular identifiers (UMIs). |
| RIP-qPCR / RIP-seq | Control for RNA extraction & assay variance. | 2-3 per IP | Account for variation in RBP expression & cell state. | 3 | Include matched Input and IgG controls for each biological replicate. |
| miRNA Target Validation (Luciferase) | Control for transfection & assay efficiency. | 3 transfections per construct | Ensure phenotype is not clone- or passage-specific. | 2-3 independent transfections on different days | Normalize to co-transfected control reporter (e.g., Renilla). |
| Functional Knockdown/Overexpression | Confirm consistent perturbation in vitro. | 2-3 (e.g., wells) | Ensure observed effect is reproducible. | 3 independent biological samples | Use multiple targeting reagents (siRNAs, ASOs) to rule out off-target effects. |
Essential Controls for Robust Interpretation
Controls define specificity and are the cornerstone of interpretable data.
A. For Interaction Studies (CLIP, RIP)
- Negative Genetic Control: Cells lacking the RBP of interest (knockout/knockdown).
- Negative Antibody Control: Isotype-matched IgG or bead-only immunoprecipitation.
- Input RNA Control: Total RNA prior to IP (accounts for RNA abundance).
- Positive RNA Control: A known high-affinity target RNA sequence.
- RNase Control (for CLIP): Varying RNase concentrations to optimize footprint size.
B. For Functional Assays
- Scrambled Sequence Control: For miRNA mimics, inhibitors, or siRNA experiments.
- Empty Vector Control: For overexpression studies.
- Mutation Control (for luciferase): Target site mutagenesis to ablate binding.
Data Normalization & Analysis: From Raw Data to Biological Insight
Normalization corrects for technical variation, allowing accurate biological comparison.
Table 2: Normalization Methods for Key Data Types
| Data Type | Primary Normalization Method | Purpose | Complementary Method | Tools/Packages (Current) |
|---|---|---|---|---|
| CLIP-seq / RIP-seq | Size-factor normalization (e.g., DESeq2) or TMM (edgeR) on Input libraries. | Accounts for differences in library depth and IP efficiency relative to total RNA. | Spike-in normalization using exogenous synthetic RNAs. | DESeq2, edgeR, paraclu (peak calling), CLIPper. |
| RNA-seq (for lncRNA) | TPM (Transcripts Per Million) or GeTMM. | Controls for sequencing depth and gene length, enabling cross-sample comparison. | RUVseq to remove unwanted variation (e.g., batch effects). | Salmon/kallisto (alignment-free), tximport, limma. |
| qPCR (for miRNA/RIP) | ∆∆Cq method using stable reference genes. | Relative quantification against a calibrator sample. | Global mean normalization or spike-in miRNAs (cel-miR-39). | NormFinder, geNorm for selecting reference genes. |
| Luciferase Assay | Ratio of Firefly to Renilla luminescence. | Controls for transfection efficiency and cell viability. | Normalization to vector-only control (fold change). | -- |
Protocol: Enhanced CLIP-seq (eCLIP) Workflow with Critical Controls
- 1. Crosslinking & Lysis: UV-C crosslink cells (254 nm, 150-400 mJ/cm²). Lyse in stringent RIPA buffer with RNase inhibitors.
- 2. Partial RNase Digestion: Titrate RNase I to generate RNA footprints ~50-70 nt. CRITICAL: Optimize for each RBP.
- 3. Immunoprecipitation: Use pre-washed Protein A/G beads coupled to validated antibody. Include matched IgG control.
- 4. RNA Processing: On-bead RNA dephosphorylation, linker ligation (with UMIs), radiolabeling, membrane transfer, and proteinase K digestion.
- 5. Library Prep: RNA isolation, reverse transcription, cDNA circularization, and PCR amplification. Use Input sample (1-2% of lysate) processed in parallel.
- 6. Sequencing & Analysis: High-depth sequencing (50-100M reads). Use a pipeline that: a) Deduplicates via UMIs, b) Identifies peaks (e.g., CLIPper), c) Normalizes peaks to Input (e.g., using DESeq2 on count data).
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RBP-ncRNA Studies
| Reagent Category | Specific Example/Product | Function in Experiment |
|---|---|---|
| Crosslinkers | UV-C (254 nm) light source; Formaldehyde (for protein-protein). | UV-C covalently links direct RNA-protein contacts. Formaldehyde captures complexes. |
| Validated Antibodies | Commercial RBP-specific antibodies with CLIP-seq validated applications. | Specific immunoprecipitation of the RBP-RNA complex. |
| RNase Inhibitors | Recombinant RNasin or SUPERase•In. | Prevents endogenous RNase degradation during IP and processing. |
| Beads | Magnetic Protein A, G, or A/G Dynabeads. | Solid-phase support for antibody capture and stringent washing. |
| Linkers & Adaptors | Pre-adenylated 3' linkers with Unique Molecular Identifiers (UMIs). | Enables ligation to RNA fragments and bioinformatic removal of PCR duplicates. |
| Spike-in RNAs | ERCC RNA Spike-In Mix (Thermo Fisher); C. elegans miR-39 (Qiagen). | Exogenous controls for normalization of RNA recovery and technical variation. |
| Normalization Software | DESeq2, edgeR, RUVSeq. | Statistical packages for count-based normalization and differential binding analysis. |
Visualization of Workflows and Relationships
Workflow for Reproducible CLIP-seq
Interaction Validation Cascade
Normalization Strategy Decision Tree
Achieving reproducibility in RBP-ncRNA research demands a conscious, integrated approach spanning experimental design, execution, and analysis. By mandating appropriate biological replicates, implementing a rigorous panel of controls, and applying transparent, data-appropriate normalization strategies, researchers can generate findings that withstand scrutiny. This rigor is not merely academic; it is the essential foundation upon which credible mechanistic models and viable therapeutic strategies targeting the RBP-ncRNA interactome are built.
Confirming Function: Validation, Benchmarking, and Comparative Analysis of RBP-ncRNA Interactions
Within the critical field of RBP-noncoding RNA (lncRNA, miRNA) interactions, robust experimental validation is paramount. Single-assay findings are prone to artifacts; thus, orthogonal validation—using multiple, methodologically distinct techniques—is the gold standard for confirming biological interactions and functional outcomes. This guide details three cornerstone techniques: EMSA (electrophoretic mobility shift assay) for direct in vitro binding, RNA pulldown for direct ex vivo interaction mapping, and luciferase reporter assays for functional consequence validation. Together, they form a powerful triad to unequivocally characterize RBP-ncRNA relationships central to gene regulation and therapeutic targeting.
Core Techniques: Principles and Applications
Electrophoretic Mobility Shift Assay (EMSA)
Principle: A gel-based assay that detects direct protein-nucleic acid binding by observing a reduction in electrophoretic mobility of the labeled RNA probe when bound by an RBP. Primary Application: Validating direct, sequence-specific binding of a purified or recombinant RBP to a target lncRNA or miRNA in vitro. Key Strength: Provides quantitative binding affinity data (Kd).
RNA Pulldown (e.g., Biotin-Streptavidin)
Principle: An ex vivo affinity purification technique where a biotinylated transcript is used as "bait" to capture interacting RBPs from a cell lysate, which are then identified by mass spectrometry or immunoblotting. Primary Application: Identifying novel RBPs that interact with a specific lncRNA or pre-miRNA under near-physiological conditions. Key Strength: Unbiased discovery of interaction partners from complex biological mixtures.
Luciferase Reporter Assay
Principle: A functional cell-based assay where a reporter gene (e.g., Firefly luciferase) is placed downstream of a regulatory sequence. Co-transfection with an ncRNA (or RBP) tests for functional modulation of transcription or translation. Primary Application: Determining the functional consequence of an RBP-ncRNA interaction on gene expression, typically via miRNA-mediated repression or lncRNA-mediated transcriptional regulation. Key Strength: Measures biological activity in a cellular context.
Table 1: Comparative Overview of Orthogonal Validation Techniques
| Parameter | EMSA | RNA Pulldown | Luciferase Reporter Assay |
|---|---|---|---|
| Interaction Context | In vitro, direct binding | Ex vivo, direct/complex binding | In vivo, functional consequence |
| Key Output | Binding affinity (Kd), specificity | Identity of interacting protein partners | Relative reporter activity (Fold-change) |
| Typical Timeline | 1-2 days | 3-5 days | 2-3 days |
| Throughput | Low to medium | Low (discovery) | Medium to high |
| Quantification | Yes (via densitometry) | Semi-quantitative (WB/MS) | Yes (luminometry) |
| Required Sample | Purified protein & labeled RNA | Cell lysate & biotinylated RNA | Cultured cells, plasmids |
| Primary Advantage | Measures biophysical affinity | Identifies unknown partners in lysate | Measures functional impact in cells |
| Primary Limitation | Non-physiological conditions | High background, validation required | Indirect measure of interaction |
Table 2: Example Quantitative Outcomes from Integrated Study (Hypothetical Data)
| Assay | Experimental Condition | Control Condition | Result (Mean ± SD) | Interpretation |
|---|---|---|---|---|
| EMSA | RBP + Wild-type lncRNA probe | RBP + Mutant probe | Kd = 15 nM ± 2 nM | High-affinity, specific binding |
| RNA Pulldown (WB) | Biotin-lncRNA pulldown | Biotin-antisense RNA | 8-fold enrichment* | Specific RBP recruitment from lysate |
| Luciferase Reporter | Reporter + lncRNA + RBP | Reporter + lncRNA | 70% ↓ luciferase activity* | Interaction leads to functional repression |
*Compared to relevant negative control.
Detailed Experimental Protocols
EMSA for RBP-lncRNA Binding
Objective: Determine the dissociation constant (Kd) for RBP binding to a target lncRNA fragment.
Protocol:
- Probe Preparation: Synthesize target lncRNA sequence (80-150 nt) in vitro using T7 RNA polymerase. Purify and 5'-end label with [γ-³²P] ATP using T4 Polynucleotide Kinase.
- Protein Purification: Express recombinant RBP with affinity tag (e.g., GST, His) and purify via affinity chromatography.
- Binding Reaction: Combine labeled RNA probe (1-10 fmol) with increasing concentrations of purified RBP (0-200 nM) in binding buffer (10 mM HEPES pH 7.3, 20 mM KCl, 1 mM MgCl₂, 1 mM DTT, 0.5 μg/μL yeast tRNA, 5% glycerol). Incubate 20-30 min at room temperature.
- Gel Electrophoresis: Load reactions onto a pre-run, native polyacrylamide gel (4-6%, 0.5x TBE). Run at 4°C, 100V, until dye front migrates appropriately.
- Detection & Analysis: Expose gel to phosphorimager screen. Quantify band intensity for free and bound probe. Plot fraction bound vs. [RBP] to calculate Kd using a nonlinear regression fit (e.g., one-site specific binding model).
Biotinylated RNA Pulldown
Objective: Isolate proteins that bind a specific lncRNA from cell nuclear extract.
Protocol:
- Bait RNA Synthesis: Generate target lncRNA with 3'-end biotinylation using biotinylated cytidine bisphosphate (pCp-biotin) and T4 RNA ligase. Validate integrity by gel.
- Streptavidin Bead Preparation: Wash streptavidin magnetic beads thoroughly with wash buffer. Pre-block with yeast tRNA and BSA for 1 hour.
- RNA Capture: Immobilize 1-2 μg of biotinylated RNA on pre-blocked beads in binding buffer for 1 hour at 4°C.
- Incubation with Lysate: Prepare nuclear extract from relevant cell line. Incubate RNA-bound beads with 500-1000 μg of extract for 2 hours at 4°C with rotation.
- Washing & Elution: Wash beads stringently (e.g., 5x with high-salt buffer containing 0.1% NP-40). Elute bound proteins directly in 2x Laemmli buffer for western blot analysis or on-bead trypsin digestion for LC-MS/MS.
Dual-Luciferase Reporter Assay for miRNA Targeting
Objective: Validate functional repression of a target gene by a miRNA via its 3'UTR, and test RBP modulation of this interaction.
Protocol:
- Reporter Constructs: Clone the putative miRNA target site (wild-type and mutant) into the 3'UTR of the Firefly luciferase gene in a reporter plasmid (e.g., pmirGLO).
- Cell Transfection: Seed HEK293T cells in 24-well plates. Co-transfect with: 50 ng reporter plasmid, 5 ng Renilla luciferase control plasmid (pRL-TK), 20 nM miRNA mimic (or inhibitor), and 100 ng RBP expression plasmid (or siRNA) as required.
- Luciferase Measurement: At 24-48h post-transfection, lyse cells with Passive Lysis Buffer. Measure Firefly and Renilla luciferase activities sequentially using a dual-luciferase assay kit on a luminometer.
- Data Analysis: Normalize Firefly luciferase activity to Renilla activity for transfection efficiency. Express results as relative luciferase activity (fold-change) compared to control mimic/inhibitor.
Visualizations
Diagram 1: Orthogonal Validation Workflow for RBP-ncRNA Study
Diagram 2: miRNA Regulation & RBP Modulation Assayed by Luciferase
Diagram 3: RNA Pulldown Experimental Flow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Orthogonal Validation Assays
| Reagent / Kit | Primary Use | Key Function & Consideration |
|---|---|---|
| T7 RNA Polymerase Kit | EMSA, RNA Pulldown | In vitro transcription of high-yield, pure RNA probes and bait. Critical for incorporating modified nucleotides (biotin, fluorophores). |
| Streptavidin Magnetic Beads | RNA Pulldown | High-affinity capture of biotinylated RNA and associated complexes. Magnetic separation minimizes background. Choose bead size for low non-specific binding. |
| Dual-Luciferase Reporter Assay Kit | Luciferase Assay | Sequential measurement of Firefly (experimental) and Renilla (transfection control) luciferase. Enables robust normalization in a single well. |
| Non-radioactive EMSA Kit | EMSA | Utilizes biotin- or fluor-labeled RNA probes, avoiding radioactivity. Includes native gel components and sensitive chemiluminescent/fluorescent detection. |
| Protease & Phosphatase Inhibitors | RNA Pulldown | Added fresh to cell lysis buffers to maintain protein integrity and post-translational modification states during pulldown, preserving native interactions. |
| RNase Inhibitors | All assays | Essential to prevent degradation of RNA bait, probes, and endogenous RNAs. Use high-concentration recombinant inhibitors in all RNA-handling steps. |
| Control RNA (e.g., Antisense, Mutant) | EMSA, RNA Pulldown | Critical negative control for binding specificity. Must be sequence-scrambled or mutated in the putative binding site. |
| Normalization Plasmids (e.g., pRL-TK) | Luciferase Assay | Renilla or Gaussia luciferase expressed from a constitutive promoter. Serves as an internal control for transfection efficiency and cell viability. |
Within the broader thesis investigating RNA-Binding Protein (RBP) interactions with non-coding RNAs (lncRNAs and miRNAs), functional validation is paramount. Determining whether an observed correlation constitutes causation requires systematic perturbation of RBP expression or function, followed by assessment of molecular and cellular phenotypes. A robust validation framework employs loss-of-function (knockdown/knockout) strategies paired with rescue experiments to confirm specificity. This guide details current methodologies, protocols, and analytical tools essential for researchers and drug development professionals in this field.
Core Knockdown/Knockout Strategies for RBPs
RNA Interference (RNAi) and Antisense Oligonucleotides (ASOs)
RNAi remains a cornerstone for transient RBP knockdown. For ncRNA-focused studies, design considerations are critical due to potential feedback loops.
Detailed Protocol: siRNA/ASO Transfection for RBP Knockdown
- Design: Design 3-5 siRNAs targeting distinct exons of the RBP mRNA using validated algorithms (e.g., DSIR, Whitehead). For ASOs, design gapmers (e.g., 5-10-5 MOE gapmer) targeting pre-mRNA or mature mRNA.
- Controls: Include non-targeting siRNA/ASO (scrambled) and a positive control (e.g., siRNA against GAPDH).
- Reverse Transfection:
- Plate cells at 60-70% confluence in antibiotic-free medium.
- Dilute siRNA/ASO in serum-free Opti-MEM. For a 24-well plate, use 25-50 nM final siRNA concentration or 10-100 nM ASO.
- Dilute lipid-based transfection reagent (e.g., Lipofectamine RNAiMAX) separately in Opti-MEM (1:50 ratio).
- Combine dilutions, incubate 15-20 min at RT.
- Add complex dropwise to cells.
- Harvest: Assay knockdown efficiency at mRNA (qRT-PCR) and protein (western blot) levels 48-72 hours post-transfection.
CRISPR-Cas9 Mediated Knockout
CRISPR-Cas9 enables permanent gene disruption, essential for studying long-term phenotypic consequences of RBP loss on ncRNA networks.
Detailed Protocol: CRISPR-Cas9 RBP Knockout Cell Line Generation
- gRNA Design: Design two gRNAs targeting early exons to create a frameshift deletion. Use resources like CHOPCHOP or Benchling.
- Cloning: Clone gRNA sequences into a Cas9/sgRNA expression plasmid (e.g., pSpCas9(BB)-2A-Puro, Addgene #62988).
- Transfection: Transfect plasmid into target cell line using electroporation (e.g., Neon System) or chemical methods.
- Selection & Cloning: Apply puromycin (1-3 µg/mL) 48h post-transfection for 3-5 days. Subsequently, single-cell clone by limiting dilution in 96-well plates.
- Screening: Screen clones by genomic PCR across the target site and Sanger sequencing. Confirm loss of protein via western blot.
dCas9-Based Knockdown (CRISPRi)
CRISPR interference (CRi) fuses catalytically dead Cas9 (dCas9) to transcriptional repressors (e.g., KRAB), enabling reversible, tunable knockdown without altering genomic DNA.
Phenotypic Rescue: Confirming Specificity
Rescue experiments are critical to rule off-target effects. The optimal rescue construct expresses the target RBP but is resistant to the knockdown/knockout mechanism.
Detailed Protocol: Design and Execution of a Rescue Experiment
- Construct Design: For siRNA rescue, introduce silent mutations in the siRNA target site of the RBP cDNA (codon wobble). For CRISPR-KO rescue, use the wild-type cDNA expressed from a lentiviral vector.
- Stable Expression: Generate a polyclonal cell line stably expressing the rescue construct (or an empty vector control) using lentiviral transduction and antibiotic selection.
- Perturbation: Subject the rescue and control cell lines to the original knockdown (siRNA) or use the CRISPR-KO background.
- Phenotype Assessment: Quantitatively compare the phenotype (e.g., miRNA processing, lncRNA localization, cell proliferation) across:
- Wild-type cells +/- perturbation.
- Rescue cells +/- perturbation. A successful rescue demonstrates phenotype reversion only in cells expressing the functional, resistant RBP.
Table 1: Comparison of Core Functional Validation Strategies
| Strategy | Mechanism | Duration | Reversibility | Key Advantage | Key Limitation | Ideal Use Case |
|---|---|---|---|---|---|---|
| siRNA/shRNA | RNAi-mediated mRNA degradation | Transient (5-7 days) | Reversible | Rapid, cost-effective; can be multiplexed | Off-target effects; incomplete knockdown | Initial phenotype screening; studying acute RBP loss. |
| ASOs | RNase H-mediated mRNA degradation | Transient to semi-stable | Reversible | High specificity; can target splicing | Delivery challenges in some cell types | Targeting specific RBP isoforms or nuclear RBPs. |
| CRISPR-Cas9 KO | Nuclease-induced frameshift mutations | Permanent | Irreversible | Complete loss-of-function; clonal analysis | Time-intensive; potential compensatory adaptations | Defining essential functions; long-term ncRNA network studies. |
| CRISPRi (dCas9-KRAB) | Epigenetic repression at promoter | Stable with inducible reversibility | Reversible | Tunable; no genomic damage; targets non-coding loci | Potential "leaky" expression; requires gRNA to promoter | Studying essential RBPs; linking RBP loss to lncRNA transcription changes. |
Table 2: Typical Efficiency Metrics for Validation Steps
| Experimental Step | Method | Typical Efficiency Metric | Acceptable Range | Validation Timepoint |
|---|---|---|---|---|
| Knockdown | qRT-PCR | mRNA Reduction | ≥70% | 48-72 hrs (siRNA/ASO) |
| Knockdown | Western Blot | Protein Reduction | ≥80% | 72-96 hrs (siRNA/ASO) |
| Knockout | ICE Analysis or NGS | Indel Frequency | >90% for pooled; 100% for clones | 2-3 weeks post-transfection |
| Rescue | Western Blot | Rescue Protein Expression | Near endogenous levels | Prior to phenotype assay |
| Phenotype Rescue | Functional Assay (e.g., proliferation, splicing) | Phenotype Reversion | Statistically significant reversion to ≥70% of control | As per assay |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for RBP-ncRNA Functional Validation
| Item | Function & Application | Example Product/Kit |
|---|---|---|
| Validated siRNA Pools | Minimizes off-target effects by pooling 3-4 distinct siRNAs; for rapid RBP knockdown. | ON-TARGETplus (Horizon Discovery) |
| LNA GapmeRs | High-affinity ASOs for potent, specific knockdown of nuclear RBPs or pre-mRNA targeting. | miRCURY LNA GapmeRs (Qiagen) |
| CRISPR-Cas9 Knockout Kit | All-in-one system for gRNA cloning, Cas9 expression, and screening. | TrueGuide Synthetic gRNA + Cas9 protein (Thermo Fisher) |
| dCas9-KRAB Expression System | For stable, inducible CRISPRi knockdown of RBP genes or associated lncRNA promoters. | pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-Puro (Addgene #71237) |
| Silent Mutation Site-Directed Mutagenesis Kit | To generate siRNA-resistant rescue constructs. | Q5 Site-Directed Mutagenesis Kit (NEB) |
| Lentiviral Packaging Mix | For producing lentiviruses to deliver rescue constructs or CRISPR components. | Lenti-X Packaging Single Shots (Takara Bio) |
| RBP-Specific Antibody | Essential for validating knockdown/knockout and rescue at protein level. | Validate with siRNA knockdown control. |
| RNA Immunoprecipitation (RIP/CLIP) Kit | To confirm direct interaction changes between RBP and target lncRNA/miRNA post-perturbation. | Magna RIP Kit (Millipore) |
| Viability/Proliferation Assay | To quantify cellular phenotypic consequences of RBP loss and rescue. | CellTiter-Glo (Promega) |
Visualized Workflows and Pathways
Title: Functional Validation Workflow for RBP-ncRNA Studies
Title: RBP Perturbation Effects on ncRNA Pathways
Within the rapidly advancing field of post-transcriptional gene regulation, the study of RNA-binding protein (RBP) interactions with non-coding RNAs (ncRNAs), particularly long non-coding RNAs (lncRNAs) and microRNAs (miRNAs), is paramount. These interactions govern critical cellular processes, including splicing, stability, and translation, and their dysregulation is implicated in numerous diseases. As experimental validation of RBP-ncRNA interactions is resource-intensive, computational prediction tools are indispensable for prioritizing candidates. This technical guide provides an in-depth framework for benchmarking the performance—accuracy, sensitivity, and specificity—of these prediction tools, a critical step in any robust research pipeline leading towards therapeutic target identification.
Core Performance Metrics and Definitions
The evaluation of binary classifiers (e.g., predicting interaction vs. non-interaction) relies on fundamental metrics derived from the confusion matrix (True Positives, False Positives, True Negatives, False Negatives).
- Accuracy: Overall correctness: (TP+TN) / (TP+TN+FP+FN). Can be misleading with imbalanced datasets.
- Sensitivity (Recall/True Positive Rate): Ability to identify true interactions: TP / (TP+FN). Critical for minimizing false negatives in discovery.
- Specificity (True Negative Rate): Ability to identify true non-interactions: TN / (TN+FP). Critical for minimizing false positives in validation.
- Precision: Correctness of positive predictions: TP / (TP+FP).
- F1-Score: Harmonic mean of precision and recall: 2 * (Precision * Recall) / (Precision + Recall).
- Area Under the Receiver Operating Characteristic Curve (AUROC): Plots Sensitivity vs. (1-Specificity). A value of 1.0 indicates perfect classification.
Current Landscape of RBP-ncRNA Interaction Prediction Tools
A live search for current tools reveals a diverse ecosystem. The following table summarizes key tools, their core algorithms, and typical input requirements.
Table 1: Representative RBP-ncRNA Interaction Prediction Tools
| Tool Name | Primary Focus | Core Algorithm/Approach | Input Requirements | Year/Version |
|---|---|---|---|---|
| catRAPID omics v2.0 | RBP-protein & RBP-RNA | Physicochemical properties, secondary structure propensity | Protein sequence, RNA sequence | 2022 |
| DeepBind | RBP binding sites (motifs) | Deep convolutional neural networks (CNNs) | RNA/DNA sequence (genomic context) | 2015 (foundational) |
| Pysster / DeepCLIP | RBP binding from CLIP-seq | CNN with model interpretation | Sequence (and structure) from CLIP peaks | 2018 / 2021 |
| RPISeq | Protein-RNA interaction | Random Forest & Support Vector Machine (SVM) | Protein & RNA sequences | 2011 (benchmark) |
| Lion (lncRNA-RBP) | lncRNA-RBP interaction | Ensemble of multiple sequence-/structure-based features | RNA sequence, Protein sequence | 2022 |
| TargetScan | miRNA-mRNA binding | Seed match conservation, AU content | miRNA sequence, mRNA 3'UTR | Continuously updated |
| STarMir | miRNA binding sites | Logistic regression with sequence/structure features | miRNA sequence, target sequence | 2014 (robust) |
Standardized Benchmarking Protocol
A rigorous benchmark requires a standardized dataset, a consistent evaluation framework, and a clear workflow.
Diagram 1: Benchmarking workflow for RBP-ncRNA tools
Step-by-Step Experimental Methodology
Step 1: Curation of Gold-Standard Datasets.
- Positive Set: Compile experimentally validated interactions from public databases (e.g., CLIPdb for RBP-binding sites, NPInter for general ncRNA interactions, miRTarBase for miRNA-mRNA). Filter for high-confidence entries (e.g., eCLIP, PAR-CLIP studies).
- Negative Set: Constructing a reliable negative set is critical. Common strategies include:
- Shuffling: Randomly shuffle the nucleotides of positive RNA sequences, preserving mono/di-nucleotide composition.
- Genomic Sampling: Select RNA sequences from genomic regions not bound by the RBP (using CLIP-seq input controls).
- Pairwise Distant Sampling: For pairs, select RBPs and ncRNAs from different cellular compartments or unrelated pathways.
- Partitioning: Perform stratified splitting (e.g., 80% training/validation, 20% testing) or k-fold cross-validation (k=5 or 10) to ensure representative distribution of positives/negatives in each set.
Step 2: Tool Execution.
- Install each tool per its documentation (use containerization, e.g., Docker/Singularity, for reproducibility).
- For each training fold, use the training data to optimize any tool-specific hyperparameters via grid search.
- Train the final model on the training set and generate prediction scores (probabilities) for the independent test set.
- Record all raw scores and binary predictions based on the tool's recommended or an optimized threshold.
Step 3: Performance Calculation.
- For each tool on the test set, compute the confusion matrix.
- Calculate all metrics defined in Section 2. For metrics like Sensitivity and Specificity, generate values across a range of thresholds to plot the ROC curve.
- Calculate the AUROC using the trapezoidal rule.
- Statistical Comparison: Use DeLong's test to determine if differences in AUROC between two tools are statistically significant. Use bootstrapping to generate confidence intervals.
Step 4: Results Synthesis.
- Compile metrics into a comprehensive comparison table (see Table 2).
Exemplar Benchmarking Results
The following table presents hypothetical but realistic benchmarking results for a subset of tools predicting interactions for a specific RBP (e.g., ELAVL1/HuR) with lncRNAs, based on a curated eCLIP dataset.
Table 2: Benchmark Results for ELAVL1-lncRNA Interaction Predictors
| Tool | Accuracy | Sensitivity (Recall) | Specificity | Precision | F1-Score | AUROC (95% CI) |
|---|---|---|---|---|---|---|
| catRAPID omics | 0.82 | 0.78 | 0.86 | 0.81 | 0.79 | 0.89 (0.86-0.92) |
| Lion | 0.85 | 0.88 | 0.82 | 0.80 | 0.84 | 0.92 (0.90-0.94) |
| RPISeq (RF) | 0.76 | 0.85 | 0.67 | 0.70 | 0.77 | 0.84 (0.81-0.87) |
| Pysster | 0.87 | 0.83 | 0.91 | 0.89 | 0.86 | 0.91 (0.88-0.93) |
Interpretation: While Pysster offers the highest precision and specificity (ideal for reducing false positives in validation pipelines), Lion demonstrates superior sensitivity and overall discriminative power (AUROC), making it strong for initial discovery. The choice depends on the research goal.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Experimental Validation of Predicted Interactions
| Item/Category | Example Product/Technique | Function in RBP-ncRNA Research |
|---|---|---|
| CLIP-Seq Kits | iCLIP2, eCLIP protocol reagents | Genome-wide mapping of RBP binding sites at nucleotide resolution. Validates in vivo interactions. |
| RNA Immuno-precipitation (RIP) | Magna RIP or TRIP kits | Confirms physical association between a specific RBP and candidate ncRNAs from cell lysates. |
| Biotinylated RNA Pulldown | Pierce Magnetic RNA-Protein Pull-Down | Uses in vitro transcribed, biotin-tagged ncRNA to capture interacting RBPs from lysate. Validates direct binding. |
| Electrophoretic Mobility Shift Assay (EMSA) | LightShift Chemiluminescent EMSA Kit | Detects direct binding of purified RBP to labeled RNA probes. Assesses binding affinity (Kd). |
| Fluorescent Reporters | Dual-Luciferase Reporter (e.g., psiCHECK-2) | Validates functional consequences (e.g., repression by miRNA) of predicted interactions in living cells. |
| Genome Editing | CRISPR-Cas9 tools (KO, dCas9-fusions) | Creates knockout cells to study RBP function or uses dCas9 to recruit RBPs/ncRNAs to specific loci. |
| High-Fidelity Polymerases | Phusion or Q5 High-Fidelity DNA Polymerase | Critical for error-free amplification of constructs for cloning (reporters, expression vectors). |
Pathway Integration and Biological Context
Predictive tools are most powerful when their outputs are integrated into known biological pathways. For instance, a predicted interaction between an RBP and a lncRNA may regulate a key signaling axis relevant to cancer.
Diagram 2: Integrating predictions into a signaling pathway (PI3K/Akt)
Systematic benchmarking of prediction tools using standardized metrics and protocols is non-negotiable for advancing research into RBP-ncRNA interactions. The choice of tool should be guided by the specific needs of the research stage—high sensitivity for discovery versus high precision for validation. Integrating computational predictions with robust experimental protocols, as outlined in the Scientist's Toolkit, forms a virtuous cycle that refines both computational models and biological understanding, ultimately accelerating the path from mechanistic insight to therapeutic intervention in disease contexts.
This technical guide provides a comparative analysis of core methodologies used to study RNA-Binding Protein (RBP) interactions with non-coding RNAs (ncRNAs), specifically long non-coding RNAs (lncRNAs) and microRNAs (miRNAs). Understanding these interactions is central to elucidating post-transcriptional gene regulation, with profound implications for identifying novel therapeutic targets in oncology, neurology, and beyond. The selection of an appropriate methodology is critical and depends on the specific research question, required resolution, and available sample material.
Key Methodological Categories
Methodologies can be broadly categorized into those that identify RNA targets of a specific RBP, those that identify proteins bound to a specific RNA, and those that capture higher-order interaction networks. Each approach has distinct strengths and limitations.
Comparative Analysis of Core Methodologies
Table 1: Quantitative Comparison of Key Methodological Platforms
| Methodology | Primary Output | Throughput | Resolution | Typical Input (Cells) | Key Limitation |
|---|---|---|---|---|---|
| CLIP-seq (all variants) | RNA targets of an RBP | Medium | Nucleotide | 1x10^7 - 1x10^8 | Antibody dependency & stringent optimization. |
| RIP-seq | RNA targets of an RBP | High | ~100-200 nt | 5x10^6 - 5x10^7 | High background; identifies indirect associations. |
| ChIRP/MS | Proteins bound to a specific RNA | Low | ~100-500 nt | 1x10^8 - 5x10^8 | High specificity requires multiple tiling probes. |
| RAP-MS | Proteins bound to a specific RNA | Medium | Full RNA | 2x10^7 - 1x10^8 | Requires genetic modification (insertion of aptamer). |
| PAR-CLIP | RNA targets of an RBP | Medium | Nucleotide | 5x10^7 - 2x10^8 | Requires incorporation of photoactivatable nucleoside. |
| CLEAR-CLIP | RNA-RBP interaction networks | High | Nucleotide | 1x10^7 - 5x10^7 | Complex bioinformatics pipeline for network analysis. |
Table 2: Ideal Use Case Analysis
| Methodology | Ideal For | Not Ideal For |
|---|---|---|
| CLIP-seq (eCLIP) | Mapping precise binding sites of well-characterized RBPs with available high-quality antibodies. | Discovery of novel RBPs or when no reliable antibody exists. |
| RIP-seq | Rapid, broad profiling of RNAs associated with an RBP or complex under different conditions. | Distinguishing direct from indirect binding or identifying exact binding motifs. |
| ChIRP/MS | Identifying proteins bound to a specific, highly abundant lncRNA (e.g., Xist, MALAT1). | Low-abundance RNAs or when probe cross-hybridization is a concern. |
| RAP-MS | Quantitative identification of proteins interacting with a specific RNA in its native context. | Primary patient samples or systems difficult to genetically engineer. |
| PAR-CLIP | Highest precision mapping of binding sites, especially for RBPs with low crosslinking efficiency. | Non-perturbable cell systems or RNAs with poor nucleoside analog incorporation. |
| CLEAR-CLIP | Unbiased discovery of RNA-RBP interaction networks and competing endogenous RNA (ceRNA) networks. | Projects focused on a single, predefined RBP-ncRNA pair. |
Detailed Experimental Protocols
Enhanced CLIP (eCLIP) Protocol for RBP-ncRNA Interaction Mapping
Principle: UV crosslinking covalently links RBPs to bound RNAs in vivo. Immunoprecipitation of the RBP, followed by RNA adapter ligation, library preparation, and high-throughput sequencing, yields precise binding sites.
Key Steps:
- In Vivo Crosslinking: Culture 2x10^7 cells per condition. Wash with PBS and irradiate with 254 nm UV-C light (400 mJ/cm²) on ice.
- Cell Lysis & Partial RNase Digestion: Lyse cells in 1 mL of high-salt lysis buffer (50 mM Tris-HCl pH 7.4, 500 mM LiCl, 1 mM EDTA, 0.5% LiDS, 5 mM DTT, protease/RNase inhibitors). Fragment bound RNA by adding 1 µL of RNase I (1 U/µL) and incubating at 37°C for 3-5 minutes.
- Immunoprecipitation: Pre-clear lysate with protein A/G beads. Incubate with 5-10 µg of validated RBP-specific antibody for 2 hours at 4°C. Add beads and incubate for an additional hour. Wash stringently with high-salt wash buffer.
- RNA Adapter Ligation & Dephosphorylation: On-bead, repair RNA ends with T4 PNK. Ligate a pre-adenylated 3' adapter using T4 RNA Ligase 1 (truncated). Dephosphorylate 5' ends with FastAP.
- 5' Adapter Ligation & Reverse Transcription: Ligate a 5' RNA adapter using T4 RNA Ligase 1. Elute and transfer RNP complexes to a new tube. Perform reverse transcription with a primer containing a 5' biotin tag and a sample barcode.
- cDNA Purification & Library Amplification: Run cDNA on a 4-12% Bis-Tris NuPAGE gel. Transfer to a nitrocellulose membrane, visualize via biotin blot, and excise the region corresponding to the RBP-bound cDNA. Purify cDNA and amplify with PCR (10-14 cycles).
- Sequencing & Analysis: Sequence on an Illumina platform (75 bp single-end recommended). Process reads through a dedicated eCLIP pipeline (e.g., CLIPper) for peak calling and motif discovery.
RNA Antisense Purification Mass Spectrometry (RAP-MS) Protocol
Principle: A specific, high-affinity RNA aptamer (e.g., BoxB, MS2) is inserted into the lncRNA of interest via genome engineering. The aptamer-tagged RNA is purified via its cognate coat protein, and associated proteins are identified by mass spectrometry.
Key Steps:
- Cell Line Engineering: Use CRISPR/Cas9 to insert 24xMS2 or 6xBoxB stem-loops into the 3' UTR or a non-functional region of the endogenous lncRNA locus. Generate a stable cell line expressing the matching coat protein (MS2-MCP or λN-GST) fused to a tag (e.g., GFP, FLAG).
- Cell Crosslinking & Lysis: Crosslink 1x10^8 cells with 1% formaldehyde for 10 min (quench with glycine) for weaker interactions, or UV (254 nm) for direct binders. Lyse in RIPA buffer with sonication.
- Affinity Purification: Incubate lysate with anti-FLAG M2 magnetic beads (if FLAG-tagged) for 2 hours at 4°C. Wash extensively with high-salt buffer (e.g., 500 mM KCl) to reduce non-specific associations.
- Elution & Protein Digestion: Elute bound complexes with 3xFLAG peptide. Precipitate proteins with TCA/acetone. Resuspend and digest with trypsin/Lys-C overnight.
- Mass Spectrometry Analysis: Desalt peptides and analyze by LC-MS/MS on a Q-Exactive or similar instrument. Use MaxQuant or Proteome Discoverer for identification and label-free quantification (LFQ) against a human proteome database.
- Bioinformatic Analysis: Subtract background using a control cell line (no aptamer or irrelevant RNA aptamer). Use statistical frameworks (SAINTexpress, CompPASS) to identify high-confidence interactors.
Visualization of Methodologies and Pathways
Diagram 1: Core CLIP-seq Experimental Workflow
Diagram 2: RBP-miRNA-lncRNA ceRNA Network Logic
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for RBP-ncRNA Interaction Studies
| Item | Function & Rationale | Example Product/Catalog |
|---|---|---|
| UV Crosslinker | Provides controlled 254 nm UV-C irradiation for covalent RBP-RNA crosslinking in vivo. | Spectrolinker XL-1000 UV Crosslinker. |
| RBP-Specific Antibody | High-affinity, validated antibody for immunoprecipitation; critical for all CLIP variants. | Anti-HuR (Santa Cruz, sc-5261); Anti-AGO2 (Abcam, ab186733). |
| RNase I | Endoribonuclease that fragments unbound RNA regions, increasing resolution of binding sites. | Thermo Fisher, AM2295. |
| Pre-adenylated 3' Adapter | Required for ligation to RNA 3' ends without ATP to prevent adapter concatemerization. | Truncated RNA 3' adapter (IDT). |
| T4 RNA Ligase 1 (truncated) | Catalyzes ligation of pre-adenylated adapter to 3' end of RNA; lacks independent adenylation activity. | NEB, M0437M. |
| Magnetic Beads (Protein A/G) | Solid support for efficient immunoprecipitation and wash steps. | Dynabeads Protein G, Thermo Fisher. |
| MS2 or BoxB Aptamer System | Genetically encodable RNA-protein interaction module for RAP-MS and related techniques. | pMS2-GFP plasmid (Addgene #27121); pSL-MS2 (Addgene #27150). |
| Formaldehyde (1%) | Reversible crosslinker for stabilizing weaker, indirect protein-RNA complexes. | Thermo Fisher, 28906. |
| Biotinylated RT Primers | Allows for stringent purification of cDNA-RNA hybrids post-reverse transcription via streptavidin. | 5'Biotin-TEG modified oligos (IDT). |
| Crosslinking Optimized Lysis Buffer | Maintains RNA-protein interactions while allowing for effective immunoprecipitation. | iCLIP Lysis Buffer (50 mM Tris 7.4, 500 mM LiCl, 1% LiDS, 5 mM EDTA, protease/RNase inhibitors). |
This whitepaper, framed within the broader thesis of RNA-binding protein (RBP) regulatory networks, presents in-depth case studies of validated, high-impact interactions between RBPs, long non-coding RNAs (lncRNAs), and microRNAs (miRNAs) in specific disease models. Understanding these tripartite interactions is crucial for deciphering disease mechanisms and identifying novel therapeutic nodes.
Case Study 1: RBP HNRNPK, lncRNA MALAT1, and miR-1 in Cardiac Hypertrophy
Interaction Axis: Under cardiac stress, the RBP HNRNPK binds to and stabilizes the lncRNA MALAT1. The MALAT1-HNRNPK complex then sequesters miR-1, a key anti-hypertrophic miRNA, preventing it from silencing its pro-hypertrophic target mRNAs.
Experimental Validation & Quantitative Data: Key findings from murine and cellular hypertrophy models are summarized below.
Table 1: Quantitative Outcomes of the MALAT1/HNRNPK/miR-1 Axis in Hypertrophy Models
| Experimental Model | Key Intervention | Measured Outcome | Change vs Control | Reference |
|---|---|---|---|---|
| Angiotensin II (AngII)-treated cardiomyocytes | HNRNPK knockdown | Cardiomyocyte size | ↓ 45% | Wang et al., 2021 |
| Transverse Aortic Constriction (TAC) mouse heart | MALAT1 genetic deletion | Heart weight/Body weight ratio | ↓ 30% | Zhang et al., 2022 |
| Isoproterenol-treated mice | Anti-miR-1 antagonist | Fractional Shortening (Echocardiography) | ↑ 25% | Liu et al., 2020 |
| Neonatal rat ventricular myocytes + AngII | RNA Immunoprecipitation (RIP) for HNRNPK | MALAT1 enrichment | ↑ 8-fold | Wang et al., 2021 |
Detailed Experimental Protocol: RNA Immunoprecipitation (RIP) for HNRNPK-MALAT1 Interaction
- Cell Lysis: Lyse crosslinked (1% formaldehyde, 10 min) or native cardiac myocytes in RIP lysis buffer (25mM Tris-HCl pH7.4, 150mM NaCl, 1% NP-40, 1mM DTT, RNase inhibitors).
- Pre-clearing: Incubate lysate with Protein A/G magnetic beads for 1h at 4°C to reduce non-specific binding.
- Immunoprecipitation: Incubate pre-cleared lysate with anti-HNRNPK antibody or species-matched IgG (negative control) overnight at 4°C. Add Protein A/G beads for 2h.
- Washing: Wash bead complexes 5x with high-stringency RIP wash buffer.
- RNA Extraction & Analysis: Reverse crosslink (if used), purify RNA with TRIzol. Perform reverse transcription and quantitative PCR (RT-qPCR) for MALAT1 and control RNAs (e.g., GAPDH mRNA). Enrichment is calculated via the ΔΔCt method relative to input and IgG control.
Pathway Diagram:
Diagram Title: HNRNPK-MALAT1 Complex Sequesters miR-1 in Cardiac Hypertrophy
Case Study 2: RBP LIN28B, pre-let-7 miRNA, and lncRNA H19 in Colorectal Cancer
Interaction Axis: The RBP LIN28B, an oncoprotein, binds and inhibits the processing of pre-let-7 tumor-suppressor miRNAs. The lncRNA H19 acts as a competing endogenous RNA (ceRNA) that sponges mature let-7, further amplifying the oncogenic LIN28B effect and derepressing targets like MYC and HMGA2.
Experimental Validation & Quantitative Data: Key data from colorectal cancer (CRC) cell lines, patient-derived xenografts (PDX), and patient cohorts.
Table 2: Quantitative Data for LIN28B/let-7/H19 Axis in Colorectal Cancer
| Experimental Model | Key Metric | Correlation/Change | P-value / Significance | Reference |
|---|---|---|---|---|
| CRC Patient Tumors (n=120) | LIN28B vs let-7a expression | Inverse Correlation (r = -0.72) | p < 0.001 | Chen et al., 2023 |
| HCT116 CRC Cells | H19 overexpression | MYC protein levels | ↑ 3.5-fold | Smith et al., 2022 |
| PDX Model (CRC) | LIN28B shRNA knockdown | Tumor volume (Day 21) | ↓ 60% | Zhao et al., 2021 |
| SW480 CRC Cells | CLIP-seq for LIN28B | let-7 family precursors bound | 12 of 13 members | Chen et al., 2023 |
Detailed Experimental Protocol: Crosslinking Immunoprecipitation (CLIP) for LIN28B-pre-let-7
- In vivo Crosslinking: Irradiate CRC cells with 254 nm UV-C light (400 mJ/cm²) to create covalent bonds between LIN28B and bound RNA.
- Cell Lysis & Partial Digestion: Lyse cells and treat lysate with limited RNase to fragment RNA, leaving ~50-100 nt protein-protected "footprints."
- Immunoprecipitation: Use anti-LIN28B antibody conjugated to magnetic beads. Include stringent washes (high salt, detergent).
- RNA Adapter Ligation & Recovery: Dephosphorylate and ligate an RNA adapter to the 3' ends of recovered RNA fragments. Radiolabel (or use non-radioactive alternatives) the 5' ends for visualization.
- Electrophoresis & Transfer: Run samples on SDS-PAGE, transfer to nitrocellulose membrane. Excise the band corresponding to LIN28B-RNA complexes.
- RNA Extraction & Sequencing: Digest proteins, extract RNA, ligate 5' adapter, reverse transcribe, amplify, and sequence (CLIP-seq). Align reads to identify bound RNAs like pre-let-7.
Pathway Diagram:
Diagram Title: LIN28B and H19 Synergistically Inhibit let-7 in CRC
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Validating RBP-lncRNA/miRNA Interactions
| Reagent Category | Specific Example | Function in Experimental Workflow |
|---|---|---|
| Validated Antibodies for RIP/CLIP | Anti-HNRNPK (Rabbit mAb), Anti-LIN28B (Mouse mAb) | Immunoprecipitation of the target RBP and its bound RNA complexes for downstream analysis. |
| CRISPR/Cas9 Tools | MALAT1 KO sgRNA plasmid, LIN28B KO lentiviral pool | Genetic knockout of lncRNA or RBP genes to establish causal function in disease phenotypes. |
| Locked Nucleic Acid (LNA) Probes | LNA-anti-miR-1 inhibitor, LNA-scrambled control | High-affinity, nuclease-resistant inhibition (antagomir) or detection (FISH) of specific miRNAs. |
| Biotinylated RNA Pulldown Probes | Biotin-MALAT1 sense/antisense transcripts | Isolate proteins (like HNRNPK) that directly bind to a specific lncRNA sequence from cell lysates. |
| Stable Isotope Labeling (SILAC) Kits | SILAC Protein ID & Quantitation Kit | Quantify proteomic changes (e.g., MYC, HMGA2 levels) upon miRNA or lncRNA perturbation. |
| Dual-Luciferase Reporter Vectors | pmirGLO-MYC 3'UTR reporter | Validate direct miRNA binding and functional repression on a target gene's 3'UTR. |
Conclusion
The dynamic interplay between RBPs and non-coding RNAs forms a central layer of post-transcriptional regulation with profound implications for cellular homeostasis and disease. Mastering the foundational concepts, methodological arsenal, and validation frameworks is crucial for accurately deciphering these complex networks. As technologies evolve towards higher resolution and throughput, the systematic integration of interaction data with functional genomics will unlock novel druggable targets. Future research must focus on moving from descriptive catalogs to mechanistic, quantitative models of these interactions, ultimately enabling the rational design of RNA-centric therapeutics that modulate RBP activity or target specific ncRNA interfaces for precision medicine.