Research Articles
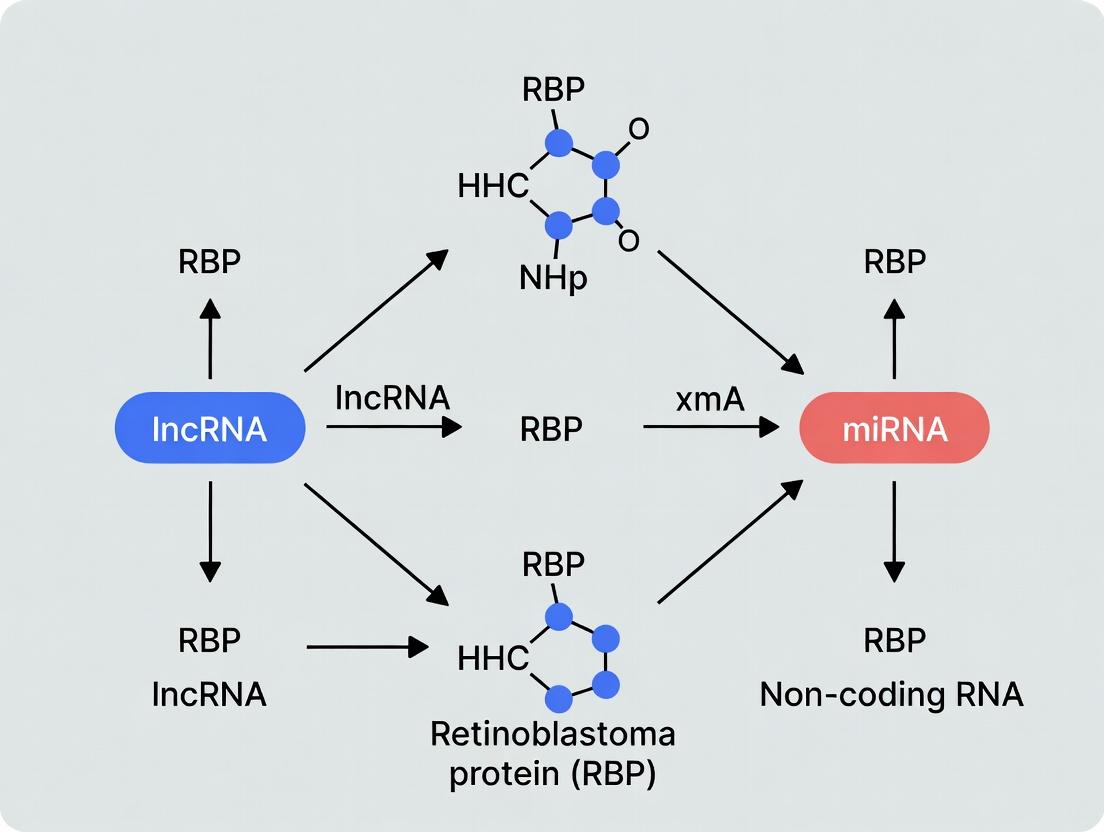
Decoding RNA Biology: A Comprehensive Guide to RBP Interactions with lncRNAs and miRNAs
This article provides a systematic overview of the intricate regulatory networks formed by RNA-binding proteins (RBPs) with non-coding RNAs, specifically long non-coding RNAs (lncRNAs) and microRNAs (miRNAs).
DESeq2 vs edgeR vs limma: Ultimate 2024 Performance Guide for Differential Expression Analysis
This comprehensive guide provides researchers, scientists, and drug development professionals with an in-depth comparison of the three leading differential expression analysis tools: DESeq2, edgeR, and limma-voom.
DESeq2 vs edgeR: A Comprehensive 2024 Guide for RNA-seq Differential Expression Analysis
This article provides a comprehensive, up-to-date comparison of DESeq2 and edgeR, the two leading R packages for differential expression analysis of RNA-seq data.
Mastering DESeq2 for RNA-Seq: A Complete Tutorial for Differential Gene Expression Analysis in Biomedical Research
This comprehensive tutorial provides researchers, scientists, and drug development professionals with a complete guide to performing differential expression analysis using DESeq2.
DESeq2 Median of Ratios Normalization: A Complete Guide for RNA-Seq Analysis in Biomedical Research
This tutorial provides a comprehensive, step-by-step guide to understanding and implementing the median of ratios normalization method in DESeq2 for RNA-seq differential expression analysis.
Beyond Accuracy: A Comprehensive Guide to Cross-Validation for RBP Binding Site Prediction
This article provides a comprehensive guide for researchers, scientists, and drug development professionals on implementing robust cross-validation (CV) strategies to assess RNA-binding protein (RBP) binding site predictors.
Benchmarking RNA Structure Prediction: A Cross-Family Performance Evaluation Guide for Researchers
Accurate RNA structure prediction is crucial for understanding gene regulation, viral function, and therapeutic target identification.
Harmony or Discord? A Comprehensive Guide to Concordance Analysis for Differential Gene Expression Tools in Biomedical Research
This article provides a systematic guide to concordance analysis for differential expression (DE) analysis tools, tailored for bioinformaticians and biomedical researchers.
Sequence vs. Structure: Which Approach Yields Higher Accuracy in RNA-Binding Protein Prediction?
This article provides a comprehensive analysis of sequence-based and structure-based methods for predicting RNA-binding proteins (RBPs), a critical task in functional genomics and drug discovery.
The RNA Folding Revolution: Traditional Algorithms vs. Machine Learning Models for Biomedical Research
This article provides a comprehensive comparison of traditional thermodynamic and kinetic algorithms with modern machine learning (ML) approaches for predicting RNA secondary and tertiary structures.